This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Andrea Foote/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 5: | Line 5: | ||
== Structure == | == Structure == | ||
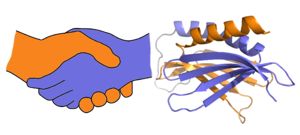
| - | <scene name='78/786627/5fgo_repeatiii/2'>PUR repeat III</scene> facilitates dimerization | + | Purα functions as a dimer composed of two intramolecular domains and one intermolecular domain. The Purα monomer contains three semi-conserved repeated amino acid sequences, named in order from N->C: PUR repeats I, II, and III. These repeats fold to form two domains: <scene name='78/786627/5fgp_intro/8'>PUR repeats I and II</scene> associating to form the I-II domain or “intramolecular domain”, while <scene name='78/786627/5fgo_repeatiii/2'>PUR repeat III</scene> facilitates dimerization through association with a repeat III from a second Purα monomer or repeat III of Purβ. Each PUR repeat is connected by flexible linker regions. |
| - | [[Image:180429 proteopedia pura figures2.jpg|thumb|right|300px| A PUR domain is analogous to a left-handed handshake. PUR repeat I-II represented from 5fgp. | + | [[Image:180429 proteopedia pura figures2.jpg|thumb|right|300px| A PUR domain is analogous to a left-handed handshake. PUR repeat I-II represented from 5fgp.]] |
| - | ]] | + | |
== Function == | == Function == | ||
| - | + | High-affinity nucleic acid-binding function is dependent on Purα dimerization. Purα forms homodimers in addition to heterodimers with Purβ. PurA is known to repress various genes including | |
<scene name='78/786627/5fgp_57and145/1'>Two aromatic residues</scene>, Y57 (repeat I) and F145 (repeat II) have been implicated in the DNA unwinding activity of PurA.<ref>PMID:26744780</ref> | <scene name='78/786627/5fgp_57and145/1'>Two aromatic residues</scene>, Y57 (repeat I) and F145 (repeat II) have been implicated in the DNA unwinding activity of PurA.<ref>PMID:26744780</ref> | ||
== Disease == | == Disease == | ||
| - | == Relevance == | ||
| - | |||
| - | == Structural highlights == | ||
| - | |||
| - | This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | ||
You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> | You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> | ||
| - | </StructureSection> | ||
| - | == Function == | ||
| - | <StructureSection load='5fgp' size='340' side='right' caption= scene='78/786627/5fgp_57and145/1'> | ||
| - | |||
| - | Anything in this section will appear adjacent to the 3D structure and will be scrollable. | ||
| - | </StructureSection> | ||
== References == | == References == | ||
<references/> | <references/> | ||
Revision as of 10:46, 3 May 2018
Purine-rich element binding protein alpha
| |||||||||||