This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Pyruvate Kinase
From Proteopedia
(Difference between revisions)
| Line 5: | Line 5: | ||
This particular protein is found in Homo sapiens and has the abbreviation PK. Pyruvate kinase belongs to the all beta proteins class and has the PK beta-barrel domain-like fold. It belongs to the PK beta-barrel domain-like superfamily and pyruvate kinase beta-barrel domain family<ref>{{website| title=SCOP: Protein: Pyruvate kinase (PK) from Human (Homo sapiens) [TaxId: 9606]|url=http://scop.berkeley.edu/data/scop.b.c.jh.b.b.d.html|}}</ref>. | This particular protein is found in Homo sapiens and has the abbreviation PK. Pyruvate kinase belongs to the all beta proteins class and has the PK beta-barrel domain-like fold. It belongs to the PK beta-barrel domain-like superfamily and pyruvate kinase beta-barrel domain family<ref>{{website| title=SCOP: Protein: Pyruvate kinase (PK) from Human (Homo sapiens) [TaxId: 9606]|url=http://scop.berkeley.edu/data/scop.b.c.jh.b.b.d.html|}}</ref>. | ||
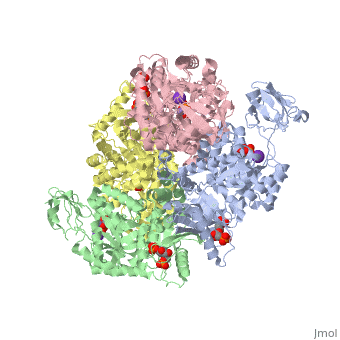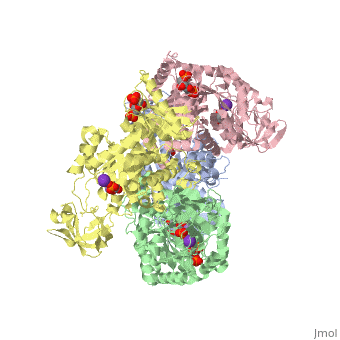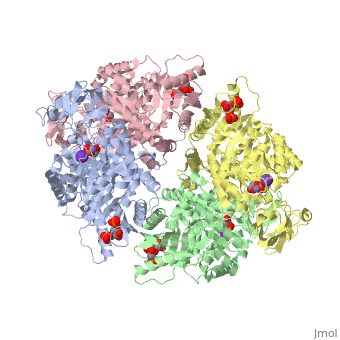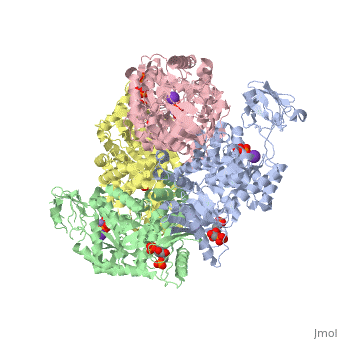
| - | Though pyruvate kinase is classified into all beta proteins, pyruvate kinase's <scene name='Keegan_Gelvoria_Sandbox_1/Secondary_structure/1'>secondary structure</scene> comprises of both alpha helices and beta sheets. In the quaternary structure of pyruvate kinase, it can be observed to have <scene name='Keegan_Gelvoria_Sandbox_1/Structure_4_domains/1'>four domains</scene> in humans. Thus, this enzyme is tetrameric with <scene name='Keegan_Gelvoria_Sandbox_1/Metal_binding_sites/1'>metal binding sites</scene> on each domain for the <scene name='Keegan_Gelvoria_Sandbox_1/Ligands/1'>K+</scene> and Mg2+ ligands to bind to. There are four types of tissue-specific isozymes: L (liver), R (red blood cells), M1 (muscle, heart, and brain), and M2 (early fetal tissue)<ref>{{website| title=SCOP: Protein: Pyruvate kinase (PK) from Human (Homo sapiens) [TaxId: 9606]|url=http://scop.berkeley.edu/data/scop.b.c.jh.b.b.d.html|}}</ref>. | + | Though pyruvate kinase is classified into all beta proteins, pyruvate kinase's <scene name='Keegan_Gelvoria_Sandbox_1/Secondary_structure/1'>secondary structure</scene> comprises of both alpha helices and beta sheets. In the quaternary structure of pyruvate kinase, it can be observed to have <scene name='Keegan_Gelvoria_Sandbox_1/Structure_4_domains/1'>four domains</scene> in humans. Thus, this enzyme is tetrameric with <scene name='Keegan_Gelvoria_Sandbox_1/Metal_binding_sites/1'>metal binding sites</scene> on each domain for the <scene name='Keegan_Gelvoria_Sandbox_1/Ligands/1'>K+</scene> and Mg2+ ligands to bind to. There are four types of tissue-specific isozymes: '''L''' (liver), '''R''' (red blood cells), '''M1''' (muscle, heart, and brain), and '''M2''' (early fetal tissue)<ref>{{website| title=SCOP: Protein: Pyruvate kinase (PK) from Human (Homo sapiens) [TaxId: 9606]|url=http://scop.berkeley.edu/data/scop.b.c.jh.b.b.d.html|}}</ref>. |
==Mechanism== | ==Mechanism== | ||
| Line 52: | Line 52: | ||
**[[3khd]] – PyK – ''Plasmodium falciparum''<br /> | **[[3khd]] – PyK – ''Plasmodium falciparum''<br /> | ||
**[[3gg8]], [[3eoe]] – PyK – ''Toxoplasma gondii''<br /> | **[[3gg8]], [[3eoe]] – PyK – ''Toxoplasma gondii''<br /> | ||
| - | **[[3bjf]], [[3bjt]], [[1t5a]], [[1zjh]] - hPyK M2 - human<br /> | + | **[[3bjf]], [[3bjt]], [[1t5a]], [[1zjh]], [[3srf]], [[3srh]] - hPyK M1/M2 - human<br /> |
| - | **[[4wj8]], [[4qg6]], [[4qg8]], [[4qg9]], [[4qgc]], [[3g2g]], [[4yj5]] - hPyK M2 (mutant)<br /> | + | **[[4wj8]], [[4qg6]], [[4qg8]], [[4qg9]], [[4qgc]], [[3g2g]], [[4yj5]] - hPyK M1/M2 (mutant)<br /> |
| - | **[[2vgb | + | **[[2vgb]] – hPyK R/L<br /> |
| - | **[[2vgf]], [[2vgg]], [[2vgi | + | **[[2vgf]], [[2vgg]], [[2vgi]] - hPyK R/L (mutant) <br /> |
**[[3ma8]], [[4drs]] – PyK – ''Cryptosporidium parvum''<br /> | **[[3ma8]], [[4drs]] – PyK – ''Cryptosporidium parvum''<br /> | ||
**[[2e28]] - PyK (mutant) – ''Geobacillus stearothermophilus''<br /> | **[[2e28]] - PyK (mutant) – ''Geobacillus stearothermophilus''<br /> | ||
**[[1f3w]] – rPyK – rabbit<br /> | **[[1f3w]] – rPyK – rabbit<br /> | ||
**[[1f3x]] - rPyK (mutant) <br /> | **[[1f3x]] - rPyK (mutant) <br /> | ||
| - | **[[ | + | **[[1pky]], [[4yng]] – EcPyK - ''Eschericia coli''<br /> |
| - | **[[ | + | **[[1e0t]], [[1e0u]] - EcPyK (mutant) <br /> |
**[[1pkm]], [[1pyk]] – PyK – cat<br /> | **[[1pkm]], [[1pyk]] – PyK – cat<br /> | ||
**[[3t05]], [[3t0t]] – SaPyK – ''Staphylococcus aureus''<br /> | **[[3t05]], [[3t0t]] – SaPyK – ''Staphylococcus aureus''<br /> | ||
**[[3qtg]] – PyK – ''Pyrobaculum aerophilum''<br /> | **[[3qtg]] – PyK – ''Pyrobaculum aerophilum''<br /> | ||
**[[4krz]] - TcPyK – ''Trypanosoma cruzei''<br /> | **[[4krz]] - TcPyK – ''Trypanosoma cruzei''<br /> | ||
| + | **[[5wrp]] - MtPyK – ''Mycobacterium tuberculosis''<br /> | ||
*Pyruvate kinase binary complex | *Pyruvate kinase binary complex | ||
| Line 77: | Line 78: | ||
**[[3is4]], [[3ktx]] – LmPyK + pyrenetetrasulfonic acid<br /> | **[[3is4]], [[3ktx]] – LmPyK + pyrenetetrasulfonic acid<br /> | ||
**[[3t07]] – SaPyK + bis-indole alkaloid<br /> | **[[3t07]] – SaPyK + bis-indole alkaloid<br /> | ||
| - | **[[4b2d]] - hPyK + fructose bisphosphate<br /> | + | **[[4b2d]] - hPyK M1/M2 + fructose bisphosphate<br /> |
| - | **[[4rpp]] - hPyK (mutant) + fructose bisphosphate<br /> | + | **[[4rpp]] - hPyK M2 (mutant) + fructose bisphosphate<br /> |
| - | **[[4fxj]] - hPyK + phenylalanine<br /> | + | **[[4fxj]] - hPyK M1/M2 + phenylalanine<br /> |
| + | **[[6gg4]] - hPyK M2 + phenylalanine<br /> | ||
| + | **[[6gg5]] - hPyK M2 + tryptophan<br /> | ||
| + | **[[6gg6]] - hPyK M2 + serine<br /> | ||
**[[5x1v]], [[5x1w]] - hPyK M2 + inhibitor<br /> | **[[5x1v]], [[5x1w]] - hPyK M2 + inhibitor<br /> | ||
**[[4rpp]] - hPyK M2 (mutant) + fructose bisphosphate<br /> | **[[4rpp]] - hPyK M2 (mutant) + fructose bisphosphate<br /> | ||
**[[4hyw]] – TbPyK + fructose bisphosphate – ''Trypanosoma brucei''<br /> | **[[4hyw]] – TbPyK + fructose bisphosphate – ''Trypanosoma brucei''<br /> | ||
| + | **[[5ws8]] - MtPyK + oxalate<br /> | ||
| - | *Pyruvate kinase | + | *Pyruvate kinase higher complex |
**[[3hqo]] – LmPyK + ATP + oxalate<br /> | **[[3hqo]] – LmPyK + ATP + oxalate<br /> | ||
| Line 90: | Line 95: | ||
**[[3qv7]] – LmPyK + acid blue 25 + Ponceau S<br /> | **[[3qv7]] – LmPyK + acid blue 25 + Ponceau S<br /> | ||
**[[3srk]] – LmPyK + saccharine + inhibitor<br /> | **[[3srk]] – LmPyK + saccharine + inhibitor<br /> | ||
| - | **[[3n25]] – rPyK + proline + Mn + pyruvate<br /> | + | **[[3n25]] – rPyK M1/M2 + proline + Mn + pyruvate<br /> |
| - | **[[2g50]] - rPyK + alanine + Mn + pyruvate<br /> | + | **[[2g50]] - rPyK M1/M2 + alanine + Mn + pyruvate<br /> |
**[[1pkn]] - rPyK + Mn + pyruvate<br /> | **[[1pkn]] - rPyK + Mn + pyruvate<br /> | ||
**[[1aqf]] – rPyK + Mg + phospholactate<br /> | **[[1aqf]] – rPyK + Mg + phospholactate<br /> | ||
**[[1a49]], [[1a5u]] - rPyK + ATP + oxalate<br /> | **[[1a49]], [[1a5u]] - rPyK + ATP + oxalate<br /> | ||
| - | **[[3me3]] – hPyK M2 + aniline derivative + fructose bisphosphate<br /> | + | **[[3me3]] – hPyK M1/M2 + aniline derivative + fructose bisphosphate<br /> |
| - | **[[4g1n]] - hPyK M2 + pyridazine derivative + oxalate<br /> | + | **[[4g1n]] - hPyK M1/M2 + pyridazine derivative + oxalate<br /> |
| - | **[[3h6o]] - hPyK M2 + pyridazine derivative + fructose bisphosphate<br /> | + | **[[3h6o]] - hPyK M1/M2 + pyridazine derivative + fructose bisphosphate<br /> |
| - | **[[3gqy]], [[3gr4]], [[3u2z]] - hPyK M2 + piperazine derivative + fructose bisphosphate<br /> | + | **[[3gqy]], [[3gr4]], [[3u2z]] - hPyK M1/M2 + piperazine derivative + fructose bisphosphate<br /> |
| - | **[[4jpg]] - hPyK M2 + pyrimidine derivative + fructose bisphosphate<br /> | + | **[[5x0i]] - hPyK M2 (mutant) + serine + fructose bisphosphate<br /> |
| - | **[[3srd]] - hPyK + oxalate + fructose bisphosphate<br /> | + | **[[5x1v]], [[5x1w]] - hPyK M2 + activator + fructose bisphosphate<br /> |
| - | **[[4fxf]] - hPyK + oxalate + ATP + fructose bisphosphate<br /> | + | **[[4jpg]] - hPyK M1/M2 + pyrimidine derivative + fructose bisphosphate<br /> |
| - | **[[4ima]], [[4ip7]] - hPyK (mutant) + citrate + adenosine + fructose bisphosphate<br /> | + | **[[3srd]] - hPyK M1/M2 + oxalate + fructose bisphosphate<br /> |
| + | **[[4fxf]] - hPyK M1/M2 + oxalate + ATP + fructose bisphosphate<br /> | ||
| + | **[[4ima]], [[4ip7]] - hPyK L (mutant) + citrate + adenosine + fructose bisphosphate<br /> | ||
**[[1a3w]] – yPyK + Mn + phosphoglycolic acid + fructose bisphosphate – yeast<br /> | **[[1a3w]] – yPyK + Mn + phosphoglycolic acid + fructose bisphosphate – yeast<br /> | ||
**[[1a3x]] - yPyK + Mn + phosphoglycolic acid<br /> | **[[1a3x]] - yPyK + Mn + phosphoglycolic acid<br /> | ||
| Line 111: | Line 118: | ||
**[[4kcw]] – TbPyK + oxalate + fructose bisphosphate <br /> | **[[4kcw]] – TbPyK + oxalate + fructose bisphosphate <br /> | ||
**[[4ks0]] – TcPyK + oxalate + fructose bisphosphate <br /> | **[[4ks0]] – TcPyK + oxalate + fructose bisphosphate <br /> | ||
| + | **[[5ws9]] – MtPyK + oxalate + AMP + ATP <br /> | ||
| + | **[[5wsa]] – MtPyK + oxalate + glucose-6-phosphate <br /> | ||
| + | **[[5wsb]], [[5wsc]] – MtPyK + oxalate + glucose-6-phosphate + AMP<br /> | ||
| + | |||
}} | }} | ||
Revision as of 10:09, 30 August 2018
| |||||||||||
3D structures of pyruvate kinase
Updated on 30-August-2018
Additional Resources
For additional information, see: Carbohydrate Metabolism
References
- ↑ Voet, Donald, Judith G. Voet, and Charlotte W. Pratt. Fundamentals of Biochemistry: Life at the Molecular Level. 3rd ed. Hoboken, NJ: John Wiley & Sons, Inc., 2008, 501-503.
- ↑ authors, The scop. "Structural Classification of Proteins". 2009. 2/26 2010. <http://scop.berkeley.edu/data/scop.b.c.jh.b.b.d.html>.
- ↑ authors, The scop. "Structural Classification of Proteins". 2009. 2/26 2010. <http://scop.berkeley.edu/data/scop.b.c.jh.b.b.d.html>.
- ↑ Robergs, Robert. "Exercise-Induced Metabolic Acidosis: Where do the Protons come from?". 2009. 2/27 2010. <http://www.sportsci.org/jour/0102/rar.htm>.
- ↑ Voet, Donald, Judith G. Voet, and Charlotte W. Pratt. Fundamentals of Biochemistry: Life at the Molecular Level. 3rd ed. Hoboken, NJ: John Wiley & Sons, Inc., 2008, 501-503.
- ↑ Voet, Donald, Judith G. Voet, and Charlotte W. Pratt. Fundamentals of Biochemistry: Life at the Molecular Level. 3rd ed. Hoboken, NJ: John Wiley & Sons, Inc., 2008, 501-503.
- ↑ Dann LG, Britton HG. Kinetics and mechanism of action of muscle pyruvate kinase. Biochem J. 1978 Jan 1;169(1):39-54. PMID:629752
- ↑ Mattevi A, Bolognesi M, Valentini G. The allosteric regulation of pyruvate kinase. FEBS Lett. 1996 Jun 24;389(1):15-9. PMID:8682196
- ↑ Jurica MS, Mesecar A, Heath PJ, Shi W, Nowak T, Stoddard BL. The allosteric regulation of pyruvate kinase by fructose-1,6-bisphosphate. Structure. 1998 Feb 15;6(2):195-210. PMID:9519410
- ↑ Oria-Hernandez J, Cabrera N, Perez-Montfort R, Ramirez-Silva L. Pyruvate kinase revisited: the activating effect of K+. J Biol Chem. 2005 Nov 11;280(45):37924-9. Epub 2005 Sep 7. PMID:16147999 doi:10.1074/jbc.M508490200
- ↑ Dann LG, Britton HG. Kinetics and mechanism of action of muscle pyruvate kinase. Biochem J. 1978 Jan 1;169(1):39-54. PMID:629752
- ↑ Oria-Hernandez J, Cabrera N, Perez-Montfort R, Ramirez-Silva L. Pyruvate kinase revisited: the activating effect of K+. J Biol Chem. 2005 Nov 11;280(45):37924-9. Epub 2005 Sep 7. PMID:16147999 doi:10.1074/jbc.M508490200
- ↑ Zanella A, Fermo E, Bianchi P, Chiarelli LR, Valentini G. Pyruvate kinase deficiency: the genotype-phenotype association. Blood Rev. 2007 Jul;21(4):217-31. Epub 2007 Mar 13. PMID:17360088 doi:10.1016/j.blre.2007.01.001
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Keegan Gelvoria, David Canner, Ann Taylor, Andrew Alexander