This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Ricardo Alberto Chiong Zevallos/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 45: | Line 45: | ||
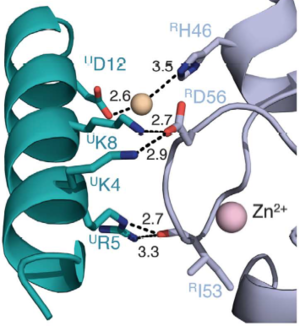
In the SPA motif of the UbcH5 (L7 residues USer94, UPro95, and UAla96), the sidechain hydroxyl of USer94 makes a hydrogen bond with the backbone carbonyl of RPro88 (from RING1b). Hydrophobic interactions between UPro95 and UAla96 with RIle53 and RPro88 help to stabilize the interaction between UbcH5 and RING1b. | In the SPA motif of the UbcH5 (L7 residues USer94, UPro95, and UAla96), the sidechain hydroxyl of USer94 makes a hydrogen bond with the backbone carbonyl of RPro88 (from RING1b). Hydrophobic interactions between UPro95 and UAla96 with RIle53 and RPro88 help to stabilize the interaction between UbcH5 and RING1b. | ||
| - | Canonical PRC1 such as BMI1/RING1b have intrinsically very low enzymatic activity compared with non-canonical PRC1, although there are only subtle differences between the structure of canonical and non-canonical complexes. Two charged helix alpha 3 residues present in a modeled BMI1 seems responsible for the low activity of BMI1/RING1b, K73 and D77 form a salt bridge that may limit efficient ubiquitin transfer. In a computational modeling, BMI1 K73 clashes sterically with Ubiquitin, which implies that K73 must move to allow Ub to bind in an activated conformation or that Ub adopts a less optimal position during monoubiquitination. Either of these events could provide an energy barrier, slowing transfer catalyzed by BMI1. The intrinsically low activity of the BMI1/RING1b is offset by a relatively favorable interaction between E3–E2-Ub and nucleosome substrate, resulting in a site-specific monoubiquitination efficient enough. The energy barrier may be responsible for increasing the fidelity of the transfer to the appropriate substrate. Also, canonical and non-canonical differ in targeting sub-units, target genomic loci, and genes expression regulation. | + | Canonical PRC1 such as BMI1/RING1b have intrinsically very low enzymatic activity compared with non-canonical PRC1, although there are only subtle differences between the structure of canonical and non-canonical complexes. Two charged helix alpha 3 residues present in a modeled BMI1 seems responsible for the low activity of BMI1/RING1b, K73 and D77 form a salt bridge that may limit efficient ubiquitin transfer. In a computational modeling, BMI1 K73 clashes sterically with Ubiquitin, which implies that K73 must move to allow Ub loaded E2 to bind in an activated conformation or that Ub adopts a less optimal position during monoubiquitination. Either of these events could provide an energy barrier, slowing transfer catalyzed by BMI1. The intrinsically low activity of the BMI1/RING1b is offset by a relatively favorable interaction between E3–E2-Ub and nucleosome substrate, resulting in a site-specific monoubiquitination efficient enough. The energy barrier may be responsible for increasing the fidelity of the transfer to the appropriate substrate. Also, canonical and non-canonical differ in targeting sub-units, target genomic loci, and genes expression regulation. |
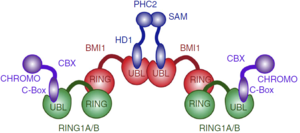
The central domain of BMI1 forms an <scene name='78/787701/5fr6_bmi1/2'>ubiquitin-like (UBL) domain</scene>, which is involved in protein-protein interactions, including interactions with the transcription factors E4F1, Zfp277 and the PLZF-RARA fusion protein. The best characterized binding partners of the UBL domain are the polyhomeotic proteins (PHC1, PHC2, PHC3). The UBL domain binds a short, 24 amino acid fragment, of PHC2 in a b-hairpin conformation. Also, UBL domain is involved in homo-oligomerization of BMI1. NMR and carbon detected NMR found that residues 30-51 are strongly conserved between PHC2, PHC1 and PHC3 suggesting that BMI1 interacts with the three members of the polyhomeotic family in a very similar manner and with similar affinities. Deletion of the corresponding motif abolished the interaction with BMI1. In the <scene name='78/787701/2na1/2'>PHC2-BMI1 complex</scene>, PHC2 residues 33–47 adopt a <scene name='78/787701/2na1_betahairpin_highlighted/1'>beta-hairpin conformation</scene> in the complex, in greenyellow. The PHC2-BMI1 interaction involves an <scene name='78/787701/2na1_bhairpin_b2_highlighted/1'>antiparallel b-sheet</scene> formed between the beta 2 strand of BMI1 UBL, in magenta, and the beta-hairpin of PHC2. The antiparallel b-sheet is stabilized by the hydrogen bonds between BMI1 Tyr163 and PHC2 Gly46. | The central domain of BMI1 forms an <scene name='78/787701/5fr6_bmi1/2'>ubiquitin-like (UBL) domain</scene>, which is involved in protein-protein interactions, including interactions with the transcription factors E4F1, Zfp277 and the PLZF-RARA fusion protein. The best characterized binding partners of the UBL domain are the polyhomeotic proteins (PHC1, PHC2, PHC3). The UBL domain binds a short, 24 amino acid fragment, of PHC2 in a b-hairpin conformation. Also, UBL domain is involved in homo-oligomerization of BMI1. NMR and carbon detected NMR found that residues 30-51 are strongly conserved between PHC2, PHC1 and PHC3 suggesting that BMI1 interacts with the three members of the polyhomeotic family in a very similar manner and with similar affinities. Deletion of the corresponding motif abolished the interaction with BMI1. In the <scene name='78/787701/2na1/2'>PHC2-BMI1 complex</scene>, PHC2 residues 33–47 adopt a <scene name='78/787701/2na1_betahairpin_highlighted/1'>beta-hairpin conformation</scene> in the complex, in greenyellow. The PHC2-BMI1 interaction involves an <scene name='78/787701/2na1_bhairpin_b2_highlighted/1'>antiparallel b-sheet</scene> formed between the beta 2 strand of BMI1 UBL, in magenta, and the beta-hairpin of PHC2. The antiparallel b-sheet is stabilized by the hydrogen bonds between BMI1 Tyr163 and PHC2 Gly46. | ||
Revision as of 23:28, 17 June 2018
| |||||||||||
References
- ↑ Jacobs JJ, Kieboom K, Marino S, DePinho RA, van Lohuizen M. The oncogene and Polycomb-group gene bmi-1 regulates cell proliferation and senescence through the ink4a locus. Nature. 1999 Jan 14;397(6715):164-8. doi: 10.1038/16476. PMID:9923679 doi:http://dx.doi.org/10.1038/16476
- ↑ Wang H, Wang L, Erdjument-Bromage H, Vidal M, Tempst P, Jones RS, Zhang Y. Role of histone H2A ubiquitination in Polycomb silencing. Nature. 2004 Oct 14;431(7010):873-8. Epub 2004 Sep 22. PMID:15386022 doi:10.1038/nature02985
- ↑ Gray F, Cho HJ, Shukla S, He S, Harris A, Boytsov B, Jaremko L, Jaremko M, Demeler B, Lawlor ER, Grembecka J, Cierpicki T. BMI1 regulates PRC1 architecture and activity through homo- and hetero-oligomerization. Nat Commun. 2016 Nov 9;7:13343. doi: 10.1038/ncomms13343. PMID:27827373 doi:http://dx.doi.org/10.1038/ncomms13343
- ↑ Bentley ML, Corn JE, Dong KC, Phung Q, Cheung TK, Cochran AG. Recognition of UbcH5c and the nucleosome by the Bmi1/Ring1b ubiquitin ligase complex. EMBO J. 2011 Jul 19. doi: 10.1038/emboj.2011.243. PMID:21772249 doi:10.1038/emboj.2011.243
- ↑ Taherbhoy AM, Huang OW, Cochran AG. BMI1-RING1B is an autoinhibited RING E3 ubiquitin ligase. Nat Commun. 2015 Jul 7;6:7621. doi: 10.1038/ncomms8621. PMID:26151332 doi:http://dx.doi.org/10.1038/ncomms8621
External resources
- Aggregated information about BMI1 protein in UniProt database
- Aggregated information about BMI1 gene in GeneCards database
https://drive.google.com/drive/folders/1l195aNuY6joOd74GKKxa-XWTRMBv_uWF?usp=sharing