This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Aminoacyl tRNA Synthetase
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
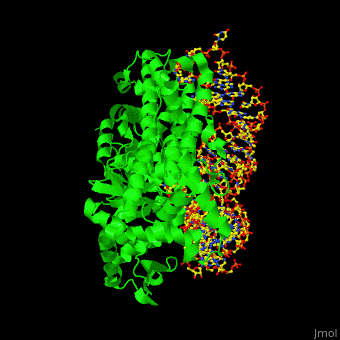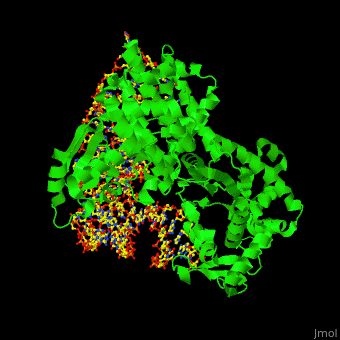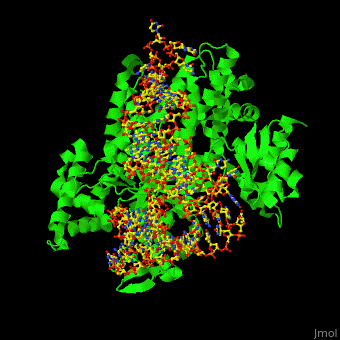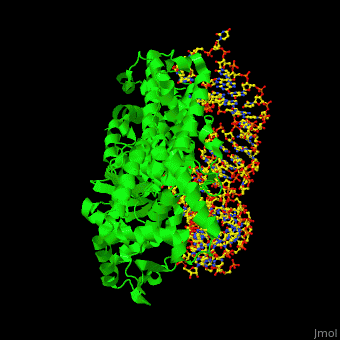
<StructureSection load='' size='350' side='right' scene='44/444597/Cv/2' caption='Arginine tRNA synthetase complex with Arg-tRNA [[1f7v]]'> | <StructureSection load='' size='350' side='right' scene='44/444597/Cv/2' caption='Arginine tRNA synthetase complex with Arg-tRNA [[1f7v]]'> | ||
| - | '''Aminoacyl tRNA synthetase''' (aaRS) catalyzes the esterification of a specific amino acid to its cognate tRNA to form an aminoacyl-tRNA. The amino acid is transferred by the ribosome from the aminoacylated-tRNA onto a growing polypeptide chain. Class I of aaRS is a monomer or dimer, it has 2 highly conserved sequence motifs and it aminoacylates at the 2’-OH of an adenosine nucleotide. Class II of aaRS is a dimer or tetramer, it has 3 highly conserved sequence motifs and it aminoacylates at the 3’-OH of an adenosine nucleotide. CP1 domain of RS edits a mischarged aa-tRNA. Some of the crystal structures are complexes of the RS with their reactant analog: amino acid-sulfamoyl adenine (aa-SA).<ref>PMID:8422978</ref>. For '''leucine-RS''' details see [[Leucyl-tRNA Synthetase]]. For '''pyrrolysyl-RS''' details see [[Pyrrolysyl-tRNA synthetase]]. | + | '''Aminoacyl tRNA synthetase''' (aaRS) catalyzes the esterification of a specific amino acid to its cognate tRNA to form an aminoacyl-tRNA. The amino acid is transferred by the ribosome from the aminoacylated-tRNA onto a growing polypeptide chain. Class I of aaRS is a monomer or dimer, it has 2 highly conserved sequence motifs and it aminoacylates at the 2’-OH of an adenosine nucleotide. Class II of aaRS is a dimer or tetramer, it has 3 highly conserved sequence motifs and it aminoacylates at the 3’-OH of an adenosine nucleotide. CP1 domain of RS edits a mischarged aa-tRNA. Some of the crystal structures are complexes of the RS with their reactant analog: amino acid-sulfamoyl adenine (aa-SA).<ref>PMID:8422978</ref>. For '''leucine-RS''' details see [[Leucyl-tRNA Synthetase]]. For '''pyrrolysyl-RS''' of '''pyrrolysine-tRNA ligase''' details see [[Pyrrolysyl-tRNA synthetase]]. |
</StructureSection> | </StructureSection> | ||
| Line 127: | Line 127: | ||
**[[1irx]] – PhLysRS<br /> | **[[1irx]] – PhLysRS<br /> | ||
**[[1bbw]] – EcLysRS<br /> | **[[1bbw]] – EcLysRS<br /> | ||
| - | **[[1krs]], [[ 1krt]] – EcLysRS anticodon-binding domain – NMR<br />[[3dsq]] – DhLysRS – ''Desulfitobacterium hafniense | + | **[[1krs]], [[ 1krt]] – EcLysRS anticodon-binding domain – NMR<br /> |
| - | + | **[[3dsq]] – DhLysRS – ''Desulfitobacterium hafniense''<br /> | |
**[[3bju]] – hLysRS residues 70-579 | **[[3bju]] – hLysRS residues 70-579 | ||
| Line 135: | Line 135: | ||
**[[3a74]] – GsLysRS + diadenosine tetraphosphate – ''Geobacillus stearothermophilus''<br /> | **[[3a74]] – GsLysRS + diadenosine tetraphosphate – ''Geobacillus stearothermophilus''<br /> | ||
**[[3e9h]], [[3e9i]] – BsLysRS + Lys-SA - ''Bacillus stearothermophilus''<br /> | **[[3e9h]], [[3e9i]] – BsLysRS + Lys-SA - ''Bacillus stearothermophilus''<br /> | ||
| - | **[[2zin]] – MmLysRS catalytic domain + Boc-Lys + ATP analog<br /> | ||
**[[1bbu]], [[1e10]], [[1lyl]] – EcLysRS + Lys<br /> | **[[1bbu]], [[1e10]], [[1lyl]] – EcLysRS + Lys<br /> | ||
**[[1e1t]] - EcLysRS + Lys-adenylate<br /> | **[[1e1t]] - EcLysRS + Lys-adenylate<br /> | ||
| Line 440: | Line 439: | ||
| - | *'''Pyrrolysyl-RS''' | + | *'''Pyrrolysyl-RS'''; domains – RNA-binding 1-101; catalytic 188-454 |
| - | **[[2e3c]] - MmPylRS catalytic domain <br /> | + | **[[2e3c]] - MmPylRS catalytic domain - ''Methanosarcina mazei''<br /> |
**[[3vqw]], [[3vqx]], [[4cs2]] - MmPylRS catalytic domain (mutant)<br /> | **[[3vqw]], [[3vqx]], [[4cs2]] - MmPylRS catalytic domain (mutant)<br /> | ||
**[[3dsq]] – DhPylRS <br /> | **[[3dsq]] – DhPylRS <br /> | ||
| Line 452: | Line 451: | ||
**[[2q7g]] - MmPylRS C-terminal+ pyrro-Lys analog+ ATP <br /> | **[[2q7g]] - MmPylRS C-terminal+ pyrro-Lys analog+ ATP <br /> | ||
**[[3vqv]] - MmPylRS catalytic domain + AMPPNP <br /> | **[[3vqv]] - MmPylRS catalytic domain + AMPPNP <br /> | ||
| - | **[[ | + | **[[4bw9]], [[4cs4]], [[5k1p]], [[5k1x]] - MmPylRS catalytic domain (mutant) + AMPPNP <br /> |
| + | **[[4bwa]] - MmPylRS catalytic domain (mutant) + adenylated norbornene <br /> | ||
**[[4ch3]], [[4ch4]], [[4ch5]], [[4ch6]] - MmPylRS catalytic domain + adenylated lysine derivative<br /> | **[[4ch3]], [[4ch4]], [[4ch5]], [[4ch6]] - MmPylRS catalytic domain + adenylated lysine derivative<br /> | ||
**[[4cs3]] - MmPylRS catalytic domain (mutant) + adenylated lysine derivative<br /> | **[[4cs3]] - MmPylRS catalytic domain (mutant) + adenylated lysine derivative<br /> | ||
| + | **[[5ud5]] - MmPylRS RNA-binding domain + Pyl-RNA<br /> | ||
| + | **[[5v6x]] - MmPylRS RNA-binding domain (mutant) + Pyl-RNA<br /> | ||
| + | **[[2zni]], [[2znj]] – DhPylRS + tRNA <br /> | ||
*''Pyrrolysyl-RS ternary complex'' | *''Pyrrolysyl-RS ternary complex'' | ||
| Line 465: | Line 468: | ||
**[[4q6g]] - MmPylRS catalytic domain + acetyl lysine + ADPNP <br /> | **[[4q6g]] - MmPylRS catalytic domain + acetyl lysine + ADPNP <br /> | ||
**[[4tqd]] - MmPylRS catalytic domain + iodo-Phe + ATP <br /> | **[[4tqd]] - MmPylRS catalytic domain + iodo-Phe + ATP <br /> | ||
| - | **[[4tqf]] - MmPylRS catalytic domain + bromothienylo-Ala + ATP <br /> | + | **[[4tqf]], [[4zib]] - MmPylRS catalytic domain + bromothienylo-Ala + ATP <br /> |
**[[2zin]] - MmPylRS catalytic domain + butoxycarbonyl lysine + ATP analog <br /> | **[[2zin]] - MmPylRS catalytic domain + butoxycarbonyl lysine + ATP analog <br /> | ||
Revision as of 07:52, 30 August 2018
| |||||||||||
3D Structures of Aminoacyl tRNA synthetase
Updated on 30-August-2018
References
- ↑ Cavarelli J, Moras D. Recognition of tRNAs by aminoacyl-tRNA synthetases. FASEB J. 1993 Jan;7(1):79-86. PMID:8422978
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Joel L. Sussman, Ann Taylor