This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2qsg
From Proteopedia
| Line 1: | Line 1: | ||
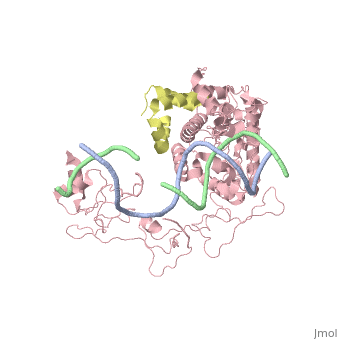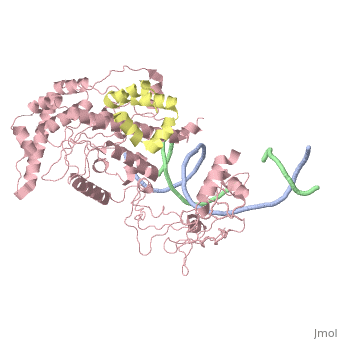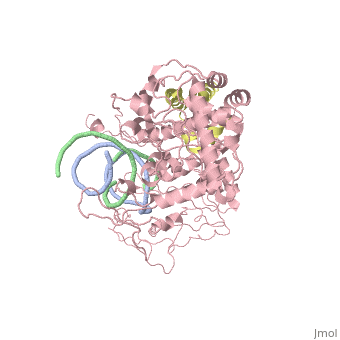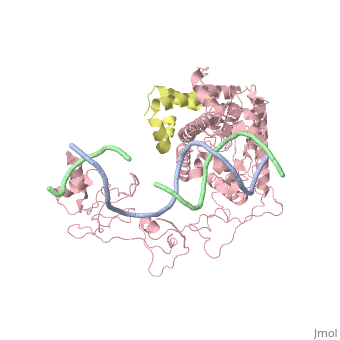
==Crystal structure of Rad4-Rad23 bound to a UV-damaged DNA== | ==Crystal structure of Rad4-Rad23 bound to a UV-damaged DNA== | ||
| - | <StructureSection load='2qsg' size='340' side='right' caption='[[2qsg]], [[Resolution|resolution]] 3.10Å' scene=''> | + | <StructureSection load='2qsg' size='340' side='right'caption='[[2qsg]], [[Resolution|resolution]] 3.10Å' scene=''> |
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[2qsg]] is a 4 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[2qsg]] is a 4 chain structure with sequence from [https://en.wikipedia.org/wiki/Atcc_18824 Atcc 18824]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2QSG OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2QSG FirstGlance]. <br> |
</td></tr><tr id='NonStdRes'><td class="sblockLbl"><b>[[Non-Standard_Residue|NonStd Res:]]</b></td><td class="sblockDat"><scene name='pdbligand=N:ANY+5-MONOPHOSPHATE+NUCLEOTIDE'>N</scene></td></tr> | </td></tr><tr id='NonStdRes'><td class="sblockLbl"><b>[[Non-Standard_Residue|NonStd Res:]]</b></td><td class="sblockDat"><scene name='pdbligand=N:ANY+5-MONOPHOSPHATE+NUCLEOTIDE'>N</scene></td></tr> | ||
| - | <tr id='related'><td class="sblockLbl"><b>[[Related_structure|Related:]]</b></td><td class="sblockDat">[[2qsf|2qsf]], [[2qsh|2qsh]]</td></tr> | + | <tr id='related'><td class="sblockLbl"><b>[[Related_structure|Related:]]</b></td><td class="sblockDat"><div style='overflow: auto; max-height: 3em;'>[[2qsf|2qsf]], [[2qsh|2qsh]]</div></td></tr> |
| - | <tr id='gene'><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">RAD4 ([ | + | <tr id='gene'><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">RAD4 ([https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=4932 ATCC 18824]), RAD23 ([https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=4932 ATCC 18824])</td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2qsg FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2qsg OCA], [https://pdbe.org/2qsg PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2qsg RCSB], [https://www.ebi.ac.uk/pdbsum/2qsg PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2qsg ProSAT]</span></td></tr> |
</table> | </table> | ||
== Function == | == Function == | ||
| - | [[ | + | [[https://www.uniprot.org/uniprot/RAD23_YEAST RAD23_YEAST]] Plays a central role both in proteasomal degradation of misfolded proteins and DNA repair. Central component of a complex required to couple deglycosylation and proteasome-mediated degradation of misfolded proteins in the endoplasmic reticulum that are retrotranslocated in the cytosol. Involved in DNA excision repair. May play a part in DNA damage recognition and/or in altering chromatin structure to allow access by damage-processing enzymes. [[https://www.uniprot.org/uniprot/RAD4_YEAST RAD4_YEAST]] Involved in nucleotide excision repair of DNA damaged with UV light, bulky adducts, or cross-linking agents. |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 39: | Line 39: | ||
</StructureSection> | </StructureSection> | ||
[[Category: Atcc 18824]] | [[Category: Atcc 18824]] | ||
| + | [[Category: Large Structures]] | ||
[[Category: Min, J H]] | [[Category: Min, J H]] | ||
[[Category: Pavletich, N P]] | [[Category: Pavletich, N P]] | ||
Revision as of 04:04, 2 July 2021
Crystal structure of Rad4-Rad23 bound to a UV-damaged DNA
| |||||||||||
Categories: Atcc 18824 | Large Structures | Min, J H | Pavletich, N P | Alpha-beta structure | Beta hairpin | Cyclobutanepyrimidine cpd dimer | Dna binding | Dna binding protein | Dna binding protein-dna complex | Dna repair | Dna-damage recognition | Mismatch dna | Nucleotide excision repair | Protein-dna complex | Transglutaminase fold | Ultraviolet uv damage | Xeroderma pigmentosum