This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
SAICAR synthetase
From Proteopedia
(Difference between revisions)
| Line 5: | Line 5: | ||
== Structural highlights == | == Structural highlights == | ||
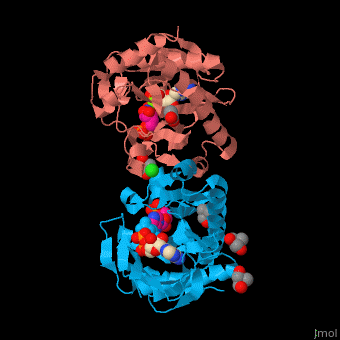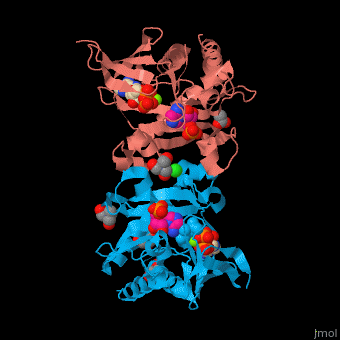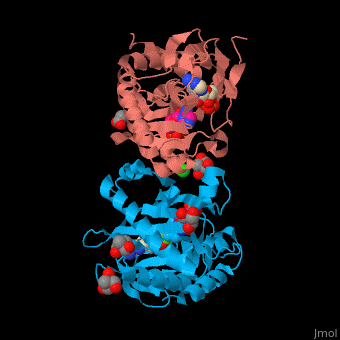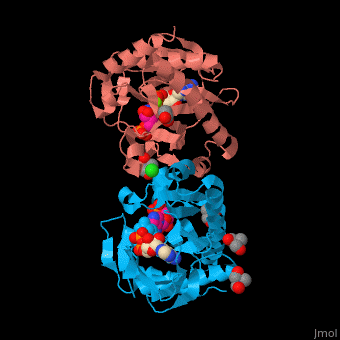
| - | SAI structure shows <scene name='74/749400/Cv/ | + | SAI structure shows <scene name='74/749400/Cv/10'>two domains</scene>. The <scene name='74/749400/Cv/11'>active site is located in a hydrophilic tunnel</scene> between the 2 domains and contains <scene name='74/749400/Cv/12'>ADP</scene>, <scene name='74/749400/Cv/13'>aspartic acid</scene> and <scene name='74/749400/Cv/14'>aminoimidazole-ribonucleotide (AIR)</scene><ref>PMID:24598753</ref>. Residues are colored according to {{Template:ColorKey_Hydrophobic}}, {{Template:ColorKey_Polar}}. Water molecules are shown as red spheres. |
| - | *<scene name='74/749400/Cv/ | + | *<scene name='74/749400/Cv/15'>Surface representation of hydrophilic tunnel</scene>. |
| - | *<scene name='74/749400/Cv/ | + | *<scene name='74/749400/Cv/16'>Mg coordination site</scene>. |
| - | *<scene name='74/749400/Cv/ | + | *<scene name='74/749400/Cv/17'>Whole active site</scene>. |
</StructureSection> | </StructureSection> | ||
Revision as of 12:43, 1 September 2019
| |||||||||||
3D structures of SAICAR synthetase
Updated on 01-September-2019
References
- ↑ Manjunath K, Jeyakanthan J, Sekar K. Catalytic pathway, substrate binding and stability in SAICAR synthetase: A structure and molecular dynamics study. J Struct Biol. 2015 Jul;191(1):22-31. doi: 10.1016/j.jsb.2015.06.006. Epub 2015, Jun 10. PMID:26072057 doi:http://dx.doi.org/10.1016/j.jsb.2015.06.006
- ↑ Wolf NM, Abad-Zapatero C, Johnson ME, Fung LW. Structures of SAICAR synthetase (PurC) from Streptococcus pneumoniae with ADP, Mg(2+), AIR and Asp. Acta Crystallogr D Biol Crystallogr. 2014 Mar;70(Pt 3):841-50. doi:, 10.1107/S139900471303366X. Epub 2014 Feb 22. PMID:24598753 doi:http://dx.doi.org/10.1107/S139900471303366X