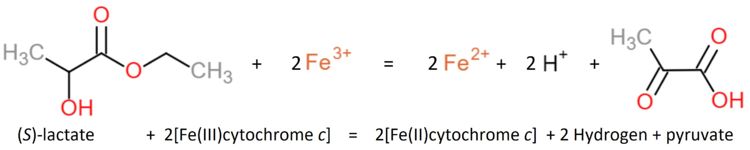This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1501
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
{{Sandbox_Reserved_ESBS}}<!-- PLEASE ADD YOUR CONTENT BELOW HERE --> | {{Sandbox_Reserved_ESBS}}<!-- PLEASE ADD YOUR CONTENT BELOW HERE --> | ||
<StructureSection load='1stp' size='340' side='right' caption='Caption for this structure' scene=''> | <StructureSection load='1stp' size='340' side='right' caption='Caption for this structure' scene=''> | ||
| - | This is a default text for your page ''''''. Click above on '''edit this page''' to modify. | ||
| - | You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> or to the article describing Jmol <ref>PMID:21638687</ref> to the rescue. | ||
| - | |||
Encoded in the ''Saccharomyces cerevisiae'' (strain ATCC 204508 / S288c) gene CYB2 the protein Flavocytochrome b(2) | Encoded in the ''Saccharomyces cerevisiae'' (strain ATCC 204508 / S288c) gene CYB2 the protein Flavocytochrome b(2) | ||
It is an Oxidoreductase categorized according to IUB as EC: 1.1.2.3. | It is an Oxidoreductase categorized according to IUB as EC: 1.1.2.3. | ||
| Line 15: | Line 12: | ||
[[Image:Reaction1.JPG|center|x150px]] | [[Image:Reaction1.JPG|center|x150px]] | ||
| - | Similar oxidants that can be used to perform the reaction in vitro are ferricyanide, phenozine methosulfate and quinone [experiment first performed 1963 by Nygaard, later in 1966 described by Symons and Burgoyne]. | + | Similar oxidants that can be used to perform the reaction in vitro are ferricyanide, phenozine methosulfate and quinone [experiment first performed 1963 by Nygaard, later in 1966 described by Symons and Burgoyne]<ref>DOI 10.1002/ijch.201300024</ref>. |
| - | The Yeasts L-Lactate Dehydrogenase can be inhibited by heavy metals, oxygen, glycerate, oxalate, malate, phenylpyruvate and fatty acids [Nygaard 1963]. | + | The Yeasts L-Lactate Dehydrogenase can be inhibited by heavy metals, oxygen, glycerate, oxalate, malate, phenylpyruvate and fatty acids [Nygaard 1963]<ref>PMID:21638687</ref>. |
The Enzyme shows a specificity for L-lactate but none for the D-isomer or -hydroxybutyrate. | The Enzyme shows a specificity for L-lactate but none for the D-isomer or -hydroxybutyrate. | ||
Revision as of 16:56, 10 January 2019
| This Sandbox is Reserved from 06/12/2018, through 30/06/2019 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1480 through Sandbox Reserved 1543. |
To get started:
More help: Help:Editing |
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644