We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Sandbox Reserved 1502
From Proteopedia
(Difference between revisions)
| Line 33: | Line 33: | ||
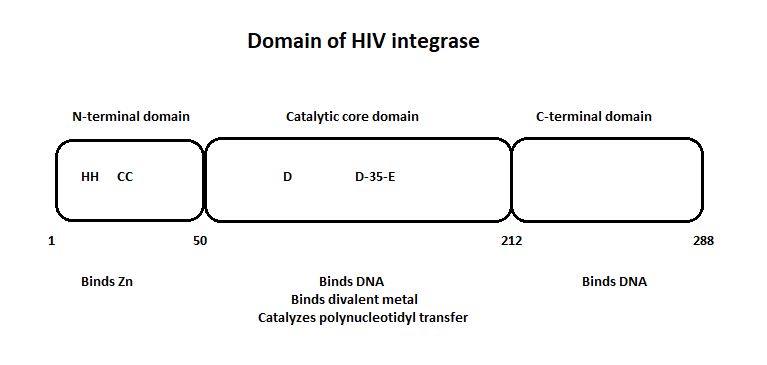
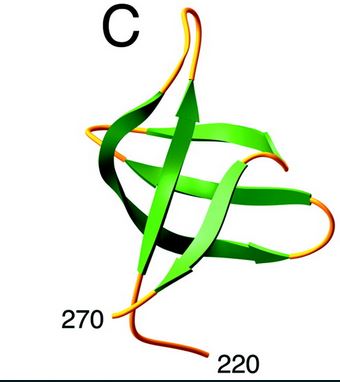
HIV-1 integrase is composed of three domains: the N-terminal (residues 1-49), the core domain (residues 50-212) and the C-terminal domain (residues 213-288). The core domain is responsible for the catalytic activity of the enzyme. It contains three acidic residues, the D,D-35-E motif which plays a key role in catalysis. The N-terminal domain includes the conserved HHCC motif, which binds zinc. The C-terminal domain is less well conserved. [http://www.jbc.org/content/276/26/23213/F2.expansion.html] | HIV-1 integrase is composed of three domains: the N-terminal (residues 1-49), the core domain (residues 50-212) and the C-terminal domain (residues 213-288). The core domain is responsible for the catalytic activity of the enzyme. It contains three acidic residues, the D,D-35-E motif which plays a key role in catalysis. The N-terminal domain includes the conserved HHCC motif, which binds zinc. The C-terminal domain is less well conserved. [http://www.jbc.org/content/276/26/23213/F2.expansion.html] | ||
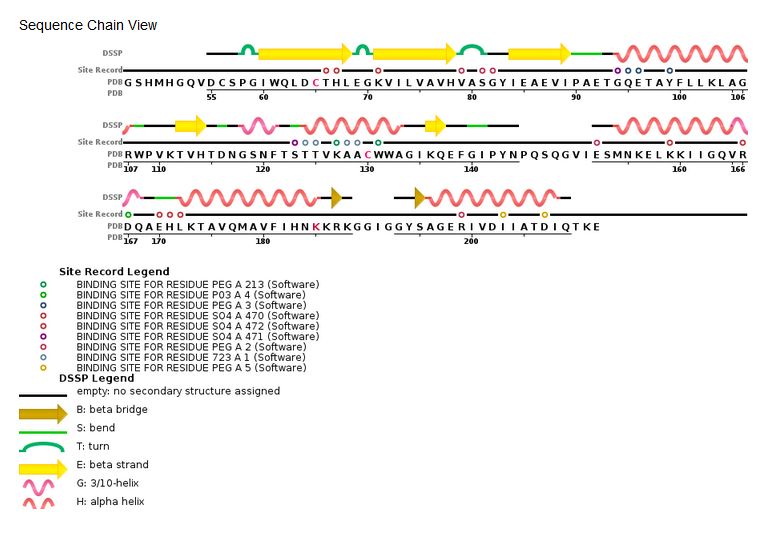
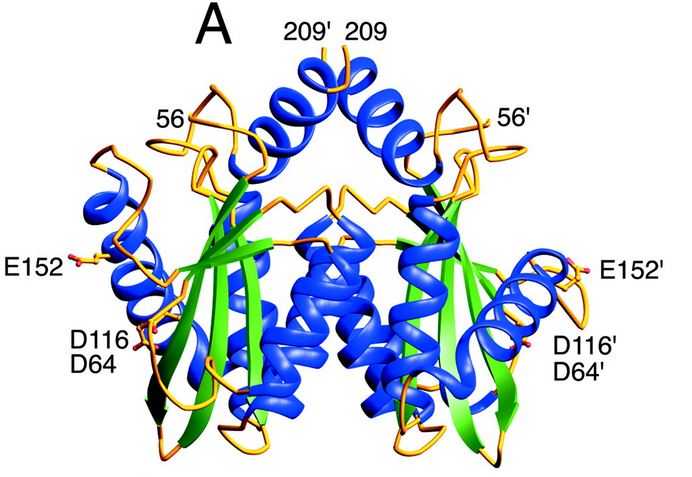
| - | ====Structure and role of the core domain in the integration==== | ||
'''The three domain of the HIV integrase:''' | '''The three domain of the HIV integrase:''' | ||
| Line 56: | Line 55: | ||
<table><tr><td colspan='2'>[[1bl3]] is a 3 chain structure with sequence from [http://en.wikipedia.org/wiki/9hiv1 9hiv1]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1BL3 OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1BL3 FirstGlance]. <br> | <table><tr><td colspan='2'>[[1bl3]] is a 3 chain structure with sequence from [http://en.wikipedia.org/wiki/9hiv1 9hiv1]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1BL3 OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1BL3 FirstGlance]. <br> | ||
</td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=MG:MAGNESIUM+ION'>MG</scene></td></tr> | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=MG:MAGNESIUM+ION'>MG</scene></td></tr> | ||
| + | |||
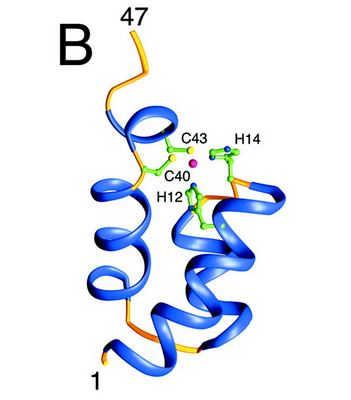
====Structure and role of the N ter domain==== | ====Structure and role of the N ter domain==== | ||
Revision as of 23:31, 10 January 2019
| This Sandbox is Reserved from 06/12/2018, through 30/06/2019 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1480 through Sandbox Reserved 1543. |
To get started:
More help: Help:Editing |
3lpt - HIV integrase
| |||||||||||