One of the CBI Molecules being studied in the University of Massachusetts Amherst Chemistry-Biology Interface Program at UMass Amherst and on display at the Molecular Playground.
Polyketides are a large and structurally diverse class of natural products produced by bacteria, fungi, and plants. They exhibit a wide variety of biological activities including antibiotic, antitumor, anticancer, among others.
In Nature, polyketides are synthesized by large multifunctional proteins called polyketide synthases (PKSs).. Among several characterized PKSs, the biosynthesis of the polyketide Core of erythromycin A, 6-deoxyerythronolide B (6-dEB), has provided the paradigm for understanding the structure and function of the PKSs that are responsible for assembling complex polyketides .
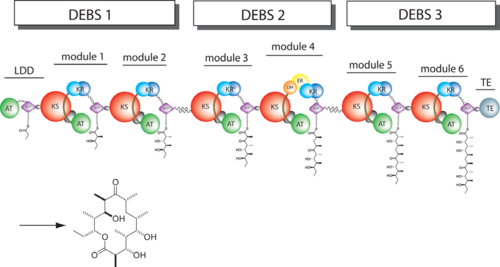
6-deoxyerythronolide B synthase (DEBS)
The 6-deoxyerythronolide B synthase (DEBS), which catalyzes the formation of 6-dEB, consists of three large subunits, DEBS1, DEBS2 and DEBS3, each containing two modules and above 300 kD in size. There are 2 domains in the N-terminal loading module, responsible for priming the synthase with a proprionate starter unit, and 26 domains in the six extender modules, Each extender module contains at least three essential domains: a ketosynthase (KS), an acyl transferase (AT) and an acyl carrier protein (ACP). In detail, the AT domain selects the appropriate carbon extender unit and transfers the units from acyl-CoA onto the phosphopantetheine arm of ACP. The KSdomain accepts the polyketide chain from the previous module and catalyzes chain elongation reaction by adding an ACP-bound extender unit through decarboxylative condensation.
After the extender unit is added, it can be further processed by optional tailoring domains, including ketoreductases (KRs), dehydratases (DHs), and enoyl reductases (ERs), to yield a hydroxyl, enoyl, or methylene group at the beta-position. Finally, the thioesterase (TE) domain that located at the C-terminus of DEBS module 6 promotes the macrocyclization event which releases the final product, 6-dEB. [1]
Ketosynthase-acyltransferase (KS-AT)
Molecular Playground banner: KS-AT The "builder" and the "gatekeeper".
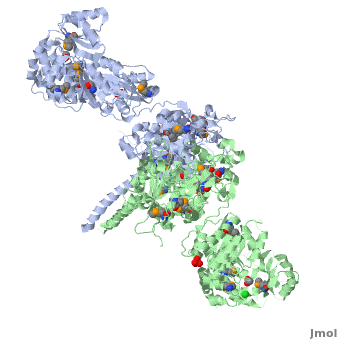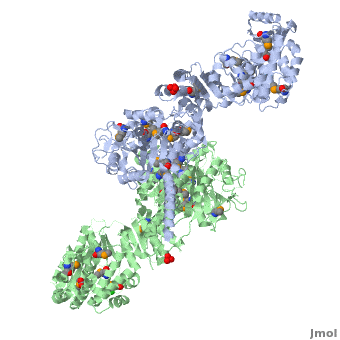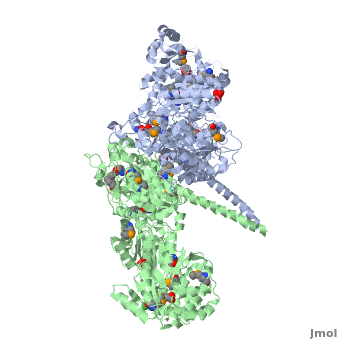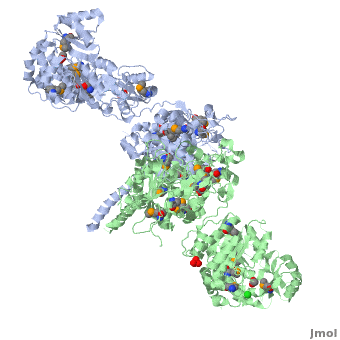
Structure of the homodimers of ketosynthase-acyltransferase didomain of module 5 from DEBS (2hg4).
The AT domain is responsible for selecting the building blocks which become incorporated into the polyketide chain, most often in the form of malonyl-CoA derivatives. Each polyketide synthase AT is exquisitely selective for one substrate type and the DEBS ATs are selective for (S)-methylmalonyl CoA. In this way it functions as the "gatekeeper", preventing erroneous building blocks from becoming incorporated into the growing chain. the methylmalonyl group is transferred by the AT to the ACP, from where it can become incorporated into the polyketide by the KS domain.
The KS accepts the "in-progress" polyketide from its upstream module, and then catalyzes the decarboxylative condensation of ACP-bound malonyl CoA derivatives with the in-progress chain. This results in an ACP-bound β-ketothioester which is processed by any other β-carbon tailoring domains present within the module.
Acyl Carrier Protein (ACP)
Molecular Playground banner: "the communicator".
(2ju1).
The ACP has no known catalytic activity. However it is central to the polyketide synthesis process by acting as a workbench upon which a round of chain extension is performed, interacting with every domain present within its module, as well as the downstream module. It accepts building blocks from AT, and then the growing chain from KS. Furthermore, it presents the extended chain to any ketoreductase (KR), dehydratase (DH), and enoyl reductase (ER) domains present, and finally passes the chain to the downstream KS or thioesterase (TE).
The heart of the ACP is the phosphopantetheine (PPant) group. An approximately 20 Å long arm which is added posttranslationally. This flexible group offers a terminal thiol through which acyl groups are bound via a thioester linkage.
Ketoreductase (KR)
The ketoreductase domain of a PKS is an optional tailoring domain that reduces a beta-ketoacyl ACP intermediate to the beta-hydroxy acyl ACP. , the co-factor for the KR domain, furnishes the hydride that is used in the reduction.
The Erythromycin KR domain
Dehydratase (DH)
Molecular Playground banner: make modification on beta-keto-acyl-ACP, "the decorator".
(3el6).
Thioesterase (TE)
Molecular Playground banner: cyclize the molecule, "the closer".
(1mo2).
TE is the terminal domain in DEBS, releasing the completed polyketide from the synthase by forming a 14-membered lactone. It is able to close such a large ring by threading it into a large inner cavity which stabilizes its conformation while esterification occurs.
3D structures of 6-deoxyerythronolide B synthase
6-deoxyerythronolide B synthase 3D structures