This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Courtney Brown/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 7: | Line 7: | ||
==Introduction== | ==Introduction== | ||
===Histones=== | ===Histones=== | ||
| - | [https://en.wikipedia.org/wiki/Histone Histones] are a family of basic, positively charged proteins that associate with DNA inside the nucleus to help condense the DNA into [https://en.wikipedia.org/wiki/Chromatin chromatin] <ref name="Histones"> Histones | Learn Science at Scitable https://www.nature.com/scitable/definition/histone-histones-57</ref>. | + | [https://en.wikipedia.org/wiki/Histone Histones] are a family of basic, positively charged proteins that associate with DNA inside the nucleus to help condense the DNA into [https://en.wikipedia.org/wiki/Chromatin chromatin] <ref name="Histones"> Histones | Learn Science at Scitable https://www.nature.com/scitable/definition/histone-histones-57</ref>. The nuclear DNA is wrapped around the histone in order to fit in the nucleus. [https://en.wikipedia.org/wiki/Nucleosome Nucleosomes] are chromatin beads made up of DNA wrapped around eight histone proteins, or a [https://en.wikipedia.org/wiki/Histone_octamer histone octamer] <ref name="Histones" />. Modifying histones is a type of [https://en.wikipedia.org/wiki/Epigenetics epigenetics], where changes are made in gene expression without altering the DNA sequence. Four different examples of modifying histones including [https://en.wikipedia.org/wiki/Histone_acetylation_and_deacetylation histone acetylation, histone deacetylation], [https://en.wikipedia.org/wiki/Histone_methylation histone methylation] and [https://en.wikipedia.org/wiki/Demethylase histone demethylation] <ref name="Histones" />. |
===Histone Deacetylases (HDACs)=== | ===Histone Deacetylases (HDACs)=== | ||
| - | ε-Amino-lysine acetylation | + | HDACs are versatile enzymes that participate in regulating non-histone proteins, the cell cycle, cell differentiation and DNA-histone interactions, which are all steps that occur during the transformation of normal cell to malignant cancer cells. They preform ε-Amino-lysine acetylation, a type of histone modification that controls the stability of proteins and biological function in eukaryotic cells <ref name="Vanninni">PMID:17721440</ref>. Histone deacetylation is the reversal process for this acetylation modification. |
| + | There are different classes of HDACs based on phylogenetic analysis: | ||
•Class I - HDACs 1-3 and 8, which are homologous to yeast Rpd3 | •Class I - HDACs 1-3 and 8, which are homologous to yeast Rpd3 | ||
| Line 20: | Line 21: | ||
•Class IV - HDAC 11 <ref name="Vanninni" />. | •Class IV - HDAC 11 <ref name="Vanninni" />. | ||
| - | HDACs 1-11 are metalloenzymes and require a zinc ion for deacetylation <ref name="Vanninni" />. | + | HDACs 1-11 are metalloenzymes and require a zinc ion for deacetylation due to their similar reaction mechanisms <ref name="Vanninni" />. |
| - | ====HDAC8==== | + | ====HDAC8==== |
[https://en.wikipedia.org/wiki/HDAC8 Histone Deacetylase 8] is an enzyme found in ''[https://en.wikipedia.org/wiki/Homo_sapiens Homo sapiens]''. HDAC8 is 388 residues long and consists of eight-stranded parallel β-sheets surrounded by 11 α-helices <ref name="Vanninni" />. HDAC8 is the only functional HDAC that is found to be a single polypeptide instead of being high-molecular-weight multi-protein complexes <ref name="Vanninni" />. HDAC8 decreases gene expression by increasing DNA-histone interactions, condensing the DNA tighter. The substrate bound to the HDAC8 includes an acetyl group, one arginine, one histidine, two lysines and MCM, a Coumarin fluorescence tag. | [https://en.wikipedia.org/wiki/HDAC8 Histone Deacetylase 8] is an enzyme found in ''[https://en.wikipedia.org/wiki/Homo_sapiens Homo sapiens]''. HDAC8 is 388 residues long and consists of eight-stranded parallel β-sheets surrounded by 11 α-helices <ref name="Vanninni" />. HDAC8 is the only functional HDAC that is found to be a single polypeptide instead of being high-molecular-weight multi-protein complexes <ref name="Vanninni" />. HDAC8 decreases gene expression by increasing DNA-histone interactions, condensing the DNA tighter. The substrate bound to the HDAC8 includes an acetyl group, one arginine, one histidine, two lysines and MCM, a Coumarin fluorescence tag. | ||
| Line 40: | Line 41: | ||
===Active Site=== | ===Active Site=== | ||
| - | + | The HDAC8 active site contains features of zinc and serine proteases. Substrates are deacetylated via a classic 2<sup>+</sup> metal ion mechanism. Zinc pentacoordinates with <scene name='81/811715/Residues_coordinating_to_zinc/1'>Asp178, His180, Asp267</scene>, the nucleophilic water and the carbonyl oxygen of the substrate<ref name="Vanninni" />. | |
| - | Substrates are deacetylated via a classic 2<sup>+</sup> metal ion mechanism. Zinc pentacoordinates with <scene name='81/811715/Residues_coordinating_to_zinc/1'>Asp178, His180, Asp267</scene>, the nucleophilic water and the carbonyl oxygen of the substrate. | + | |
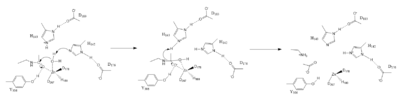
The Zinc ion <scene name='81/811715/Zn_activating_water/1'>activates the water</scene> by withdrawing electron density from the water due to the positive charge, making the water more acidic, therefore a better nucleophile. The Zinc ion also coordinates to the carbonyl oxygen of the acetyl group on the lysine, polarizing the carbonyl carbon, making it more electrophilic. <scene name='81/811715/His_142_and_143/1'>His142</scene> deprotinates the water, the first step of the deacetylation (Figure 1). His 142 (stabilized by Asp 176) is closer to the water molecule than His 143 (stabilized by Asp 183), which was also thought to deprotinate the water. However, a mutation done to His 143 only reduced activity, not abolished it, showing it is important but not crucial. His 143 is instead thought to orient the substrate<ref name="Vanninni" />. | The Zinc ion <scene name='81/811715/Zn_activating_water/1'>activates the water</scene> by withdrawing electron density from the water due to the positive charge, making the water more acidic, therefore a better nucleophile. The Zinc ion also coordinates to the carbonyl oxygen of the acetyl group on the lysine, polarizing the carbonyl carbon, making it more electrophilic. <scene name='81/811715/His_142_and_143/1'>His142</scene> deprotinates the water, the first step of the deacetylation (Figure 1). His 142 (stabilized by Asp 176) is closer to the water molecule than His 143 (stabilized by Asp 183), which was also thought to deprotinate the water. However, a mutation done to His 143 only reduced activity, not abolished it, showing it is important but not crucial. His 143 is instead thought to orient the substrate<ref name="Vanninni" />. | ||
| - | The attack by the deprotinated water forces the carbonyl carbon into a tetrehedral transition state. <scene name='81/811715/Tyr306/4'>Tyr306</scene> acts as the oxyanion hole, stabilizing this transition state via its protruding -OH group hydrogen-bonding with the negatively charged oxygen. The transition state collapses, and His 143 simultaneously gets deprotinated by the amide of the leaving acetyl group in order to weaken the scissle bond. Breaking this scissle bond results in a neutral lysine and a carboxylic acid, among other products. Body pH is around 7 typically and the pKa of carboxylic acids is around 2; the hydrogen from the COOH group is lost and picked up by the lysine, providing the hydrogen necessary for lysine to become acidic, per its nature. | + | The attack by the deprotinated water forces the carbonyl carbon into a tetrehedral transition state. <scene name='81/811715/Tyr306/4'>Tyr306</scene> acts as the oxyanion hole, stabilizing this transition state via its protruding -OH group hydrogen-bonding with the negatively charged oxygen. The transition state collapses, and His 143 simultaneously gets deprotinated by the amide of the leaving acetyl group in order to weaken the scissle bond. Breaking this scissle bond results in a neutral lysine and a carboxylic acid, among other products. Body pH is around 7 typically and the pKa of carboxylic acids is around 2; the hydrogen from the COOH group is lost and picked up by the lysine, providing the hydrogen necessary for lysine to become acidic, per its nature<ref name="Vanninni" />. |
==Disease== | ==Disease== | ||
Revision as of 22:25, 7 April 2019
The Human Histone H3/K9 Deacetylase, HDAC8
| |||||||||||