This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Courtney Brown/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 35: | Line 35: | ||
The ligand is also held in place via hydrophobic <scene name='81/811715/Hydrophobic_interactions/3'>interactions</scene> between Phe152, the nonpolar chain of the acetylated lysine, and Phe208. These hydrophobic stacking interactions keep other molecules out of the active site. Gly151 also hydrogen-bonds to a backbone carbonyl oxygen, further helping the substrate stay in place<ref name="Vanninni">PMID:17721440</ref>. | The ligand is also held in place via hydrophobic <scene name='81/811715/Hydrophobic_interactions/3'>interactions</scene> between Phe152, the nonpolar chain of the acetylated lysine, and Phe208. These hydrophobic stacking interactions keep other molecules out of the active site. Gly151 also hydrogen-bonds to a backbone carbonyl oxygen, further helping the substrate stay in place<ref name="Vanninni">PMID:17721440</ref>. | ||
| - | ===Inhibitor=== | ||
===Potassium Binding Site=== | ===Potassium Binding Site=== | ||
| Line 49: | Line 48: | ||
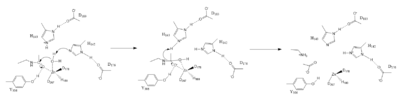
The attack by the deprotinated water forces the carbonyl carbon into a tetrehedral transition state. <scene name='81/811715/Tyr306/4'>Tyr306</scene> acts as the oxyanion hole, stabilizing this transition state via its protruding -OH group hydrogen-bonding with the negatively charged oxygen. The transition state collapses, and His 143 simultaneously gets deprotinated by the amide of the leaving acetyl group in order to weaken the scissle bond. Breaking this scissle bond results in a neutral lysine and a carboxylic acid, among other products. Body pH is around 7 typically and the pKa of carboxylic acids is around 2; the hydrogen from the COOH group is lost and picked up by the lysine, providing the hydrogen necessary for lysine to become acidic, per its nature<ref name="Vanninni" />. | The attack by the deprotinated water forces the carbonyl carbon into a tetrehedral transition state. <scene name='81/811715/Tyr306/4'>Tyr306</scene> acts as the oxyanion hole, stabilizing this transition state via its protruding -OH group hydrogen-bonding with the negatively charged oxygen. The transition state collapses, and His 143 simultaneously gets deprotinated by the amide of the leaving acetyl group in order to weaken the scissle bond. Breaking this scissle bond results in a neutral lysine and a carboxylic acid, among other products. Body pH is around 7 typically and the pKa of carboxylic acids is around 2; the hydrogen from the COOH group is lost and picked up by the lysine, providing the hydrogen necessary for lysine to become acidic, per its nature<ref name="Vanninni" />. | ||
| - | == | + | ==Inhibition== |
| - | [https://friedreichsataxianews.com/friedreichs-ataxia-experimental-treatments/histone-deacetylase-inhibitors/ HDACis] | + | ===HDACis=== |
| + | HDAC inhibitors ([https://friedreichsataxianews.com/friedreichs-ataxia-experimental-treatments/histone-deacetylase-inhibitors/ HDACis]) are a class of compounds that deactivate histone deacetylases by promoting the acetylation of specific lysine residues on histones. While a variety of HDACis are used to target different HDACs, common structural motifs include a zinc-binding moiety (ZBM) in the catalytic pocket opposite of the capping group, and a straight chain alkyl, vinyl, or aryl linker connecting the two, usually a hydrophobic chain of six carbons. HDACis bind and deactivate the HDACs through the amino acid sequence of the rim surrounding the catalytic site of the different HDACs (GREEN LINK – HDAC8 with hydroxamic acid bound). The capping group is generally hydrophobic and interacts with the rim amino acids of the HDAC. Binding occurs via interaction between conserved sections of HDAC active sites and the alkyl, vinyl, or aryl functional groups of the HDACi. The zinc binding moiety catalyzes hydrolysis of the acetyl-lysine bond at the bottom of the catalytic pocket, allowing for deactivation of the HDAC (GREEN LINK – HDAC CATALYTIC SITE WITH WITH INHIBITOR). | ||
| + | |||
| + | ===Clinical Application=== | ||
| + | A variety of HDACis are currently being tested in regards to cancer treatment. Excess amounts of HDACs have been found in tumor tissue compared to healthy tissue, implying that high amounts of HDAC are reacting with and deacetylating the p53 gene of which the HDAC is derived. The p53 gene is a tumor suppressor that when deacetylated is rendered inactive due to decreased transcriptional activity and upregulation of oncogenes. The uncontrolled growth of tumors can manifest and lead to cancer. In the past years, a variety of HDACis have been synthesized and tested for development as potential anticancer agents. The variety of HDACis has been generated by modification of the zinc binding moiety or the capping group. Modifications to the rim of amino acids on the catalytic site of HDACs have also been modified to allow for broader specificity of HDACis that can bind. Current types of HDACis include hydroxamic acids, cyclic peptides, electrophilic ketones, short chain fatty acids, benzamides, boronic acid based compounds, benzofuranone and sulfonamide. Of these, two HDACis are currently in clinical trials as cancer drugs, known as vorinostat and romidepsin. | ||
| + | |||
== References == | == References == | ||
Revision as of 03:16, 8 April 2019
The Human Histone H3/K9 Deacetylase, HDAC8
| |||||||||||