This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Nicholas Bantz/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
== Introduction == | == Introduction == | ||
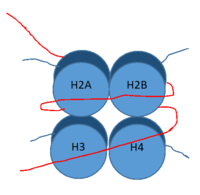
[[Image:Histone.png|200 px|right|thumb|Figure 1: DNA (red) wrapped around histone proteins with histone tails (blue)]] | [[Image:Histone.png|200 px|right|thumb|Figure 1: DNA (red) wrapped around histone proteins with histone tails (blue)]] | ||
| - | <scene name='81/811090/Overall_lsd-1/1'>LSD-1</scene>, human lysine-specific demethylase 1, is an enzyme that affects the ability of DNA to associate with [https://en.wikipedia.org/wiki/Histone histone proteins]. Histone proteins are positively charged proteins that act as spools for negatively charged DNA to wrap around for storage | + | <scene name='81/811090/Overall_lsd-1/1'>LSD-1</scene>, human lysine-specific demethylase 1, is an enzyme that affects the ability of DNA to associate with [https://en.wikipedia.org/wiki/Histone histone proteins]. Histone proteins are positively charged proteins that act as spools for negatively charged DNA to wrap around for storage in the nucleus (Figure 1). When DNA is tightly condensed it forms into nucleosomes which consist of 8 histone core proteins (2 H2A, 2 H2B, 2 H3, 2 H4) with DNA tightly coiled around them. This tightly coiled DNA is known as [https://en.wikipedia.org/wiki/Heterochromatin heterochromatin], which is inaccessible to transcription factors and RNA polymerase. This can be reversed by modifications to histone protein structure that cause the DNA to relax and form [https://en.wikipedia.org/wiki/Euchromatin euchromatin], which allows for RNA polymerase and other transcription factors to properly execute transcription. One key histone modification is the [https://en.wikipedia.org/wiki/Demethylase demethylation] of lysine residues. Before 2004, it was believed that methylation of histone tails was stable and irreversible. In 2004, it was discovered that histone tails can also be demethylated by demethylase enzymes such as LSD-1 <ref name="Shi">doi: 10.1016/j.cell.2004.12.012</ref>. LSD-1 specifically demethylates mono- or di-methylated lysine substrates at Lys4 or Lys9 in the tail of histone H3. Demethylation of these lysine residues is commonly associated with transcriptional activation, but it also has the ability to silence genes depending on the residue being demethylated, the cofactors present, and the environment in which the demethylation occurs. LSD-1 is among the most well-known demethylases and has been studied since its instrumental discovery in 2004 <ref name="Shi"/>. |
== Structure == | == Structure == | ||
=== Tower Domain === | === Tower Domain === | ||
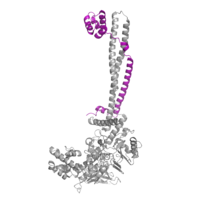
| - | [[Image:COREST.png|200 px|left|thumb|Figure 2: CoRest complex (purple) bound to LSD1 at the Tower domain.]] | + | [[Image:COREST.png|200 px|left|thumb|Figure 2: CoRest complex (purple) bound to LSD1 (PDB: 2h94) at the Tower domain.]] |
| - | The <scene name='81/811088/Towerdomain/2'>tower domain</scene> is a 100 residue protrusion off of the main protein body of LSD-1, comprised of 2 [https://en.wikipedia.org/wiki/Alpha_helix-helices 𝛂-helices]. The longer helix, T𝛂A, is an LSD-1 specific element that has not been found in any other oxidase proteins <ref name="Stavropolous">doi: 10.1038/nsmb1113</ref>. The shorter helix, T𝛂B, is positioned near the active site of the oxidase domain. | + | The <scene name='81/811088/Towerdomain/2'>tower domain</scene> is a 100 residue protrusion off of the main protein body of LSD-1, comprised of 2 right-handed [https://en.wikipedia.org/wiki/Alpha_helix-helices 𝛂-helices]. The longer helix, T𝛂A, is an LSD-1 specific element that has not been found in any other oxidase proteins <ref name="Stavropolous">doi: 10.1038/nsmb1113</ref>. The shorter helix, T𝛂B, is positioned near the active site of the oxidase domain. T𝛂B connects directly to helix 𝛂D of the oxidase domain through a highly conserved connector loop. The exact function of the tower domain is not known, but it is proposed to regulate the size of the active site chamber through this <scene name='81/811090/Tb-dinteraction/1'>TαB-αD interaction</scene>. The T𝛂B-𝛂D interaction is responsible for the proper positioning of <scene name='81/811090/Phe538-tyr761interaction/1'>Phe538</scene>, a side chain of 𝛂D that is located in the catalytic chamber, for proper recognition and binding of the substrate lysine through hydrophobic interactions. In addition, the T𝛂B-𝛂D interaction positions 𝛂D in the correct manner to provide [https://en.wikipedia.org/wiki/Hydrogen_bond hydrogen bonding] to <scene name='81/811090/Phe538-tyr761interaction/1'>Tyr761</scene>. Tyr761 is positioned in the catalytic chamber next to the FAD cofactor and aids in the binding of the lysine substrate <ref name="Stavropolous"/>. Therefore, the base of the tower domain forms a direct connection to the oxidase domain and plays a crucial role in the shape and catalytic activity of the active site. In fact, removing the tower domain via a mutation resulted in a drastic decrease in catalytic efficiency <ref name="Stavropolous"/>. The tower domain has also been found to interact with other proteins and complexes, such as CoREST (Figure 2), as a molecular lever to allosterically regulate the catalytic activity of the active site <ref name="Yang">doi: 10.1016/j.molcel.2006.07.012</ref>. Overall, the exact function of the tower domain has not yet been fully determined, but it is known to be vital to the catalytic activity of LSD-1. |
| Line 17: | Line 17: | ||
=== Oxidase Domain === | === Oxidase Domain === | ||
| - | The <scene name='81/811088/Oxidasedomain/3'>oxidase domain</scene> | + | The <scene name='81/811088/Oxidasedomain/3'>oxidase domain</scene> houses the catalytic site of LSD-1. The domain has two distinct regions: one non-covalently binds the FAD cofactor and the other acts in both the binding and recognition of the substrate lysine on a histone tail(H3)<ref name="Stavropolous"/>. The active site cavity is placed within the substrate-binding subunit of the oxidase domain and is unique due to its great size. In relation to other FAD-dependent oxidases, LSD-1 has a very large active site cavity that is 15 Å deep and 25 Å at its widest opening <ref name="Stavropolous"/>. In comparison, [https://en.wikipedia.org/wiki/Polyamine_oxidase polyamine oxidase], another FAD-dependent oxidase, has a catalytic chamber roughly 30 Å long but only a few angstroms wide <ref name=”Binda”>PMID:11258887</ref>. The relatively large size of the LSD-1 active site cavity suggests that other residues, in addition to the substrate lysine, enter into the active site during catalysis. These additional residues could participate in substrate recognition and may contribute to the enzyme’s specificity for H3K4 and H3K9. |
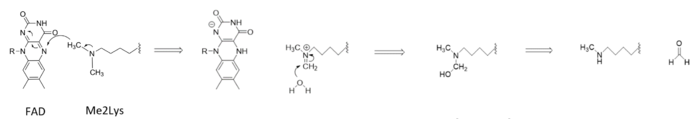
====Active Site and FAD Cofactor==== | ====Active Site and FAD Cofactor==== | ||
Revision as of 18:55, 16 April 2019
LSD-1: Human lysine-specific demethylase 1
| |||||||||||
Student Contributors
- Nicholas Bantz
- Cody Carley
- Michael Thomas