User:Caitlin Marie Gaich/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 12: | Line 12: | ||
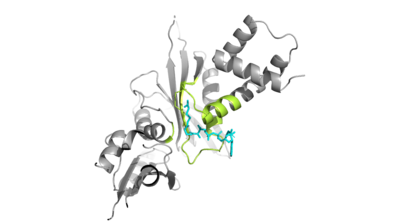
The HAT1 and HAT2 interface is stabilized by hydrogen bonds, <scene name='81/811717/Salt_bridges/1'>salt bridges</scene>, and hydrophobic interactions. Most of these interactions are located in <scene name='81/811717/Lp1/2'>LP1</scene> of the HAT1 domain, which forms a well-ordered helix. This LP1 helix is thought to be important for the heterodimer formation as the deletion of LP1 abolished the interaction between HAT1 and HAT2. This suggests that there may be another protein involved such as the N terminus tail of H4 acting as a linker protein interacting with the complex interface. The three major areas where hydrogen bonds are present aids in this complex formation. The side chain atoms of <scene name='81/811717/Tyr199_asp308_ala202/5'>Tyr199 and Asp308</scene> with the main chain nitrogen of Ala202 in HAT1. The side chain of <scene name='81/811717/Lys211phe205_and_leu288arg282/7'>Lys211 and Arg282</scene> makes hydrogen bonds with Leu288 and Phe205 respectively. The last area of hydrogen bonds between HAT1 and HAT is found between <scene name='81/811717/Serine_hydrogen_bonds/3'>Ser263 and Asp 206</scene>. The <scene name='81/811717/Hydrophobic_core/2'>hydrophobic core</scene> at the interface of the complex appears to be critical for the complex formation. This core consists of aromatic amino acids from HAT1 and leucine amino acids from HAT2, however it does not form any obvious ring stacking. | The HAT1 and HAT2 interface is stabilized by hydrogen bonds, <scene name='81/811717/Salt_bridges/1'>salt bridges</scene>, and hydrophobic interactions. Most of these interactions are located in <scene name='81/811717/Lp1/2'>LP1</scene> of the HAT1 domain, which forms a well-ordered helix. This LP1 helix is thought to be important for the heterodimer formation as the deletion of LP1 abolished the interaction between HAT1 and HAT2. This suggests that there may be another protein involved such as the N terminus tail of H4 acting as a linker protein interacting with the complex interface. The three major areas where hydrogen bonds are present aids in this complex formation. The side chain atoms of <scene name='81/811717/Tyr199_asp308_ala202/5'>Tyr199 and Asp308</scene> with the main chain nitrogen of Ala202 in HAT1. The side chain of <scene name='81/811717/Lys211phe205_and_leu288arg282/7'>Lys211 and Arg282</scene> makes hydrogen bonds with Leu288 and Phe205 respectively. The last area of hydrogen bonds between HAT1 and HAT is found between <scene name='81/811717/Serine_hydrogen_bonds/3'>Ser263 and Asp 206</scene>. The <scene name='81/811717/Hydrophobic_core/2'>hydrophobic core</scene> at the interface of the complex appears to be critical for the complex formation. This core consists of aromatic amino acids from HAT1 and leucine amino acids from HAT2, however it does not form any obvious ring stacking. | ||
| - | Once the complex has formed, histone 4 and AcetylCoA can begin to interact. The N-terminal segment of H4 that binds with HAT1/HAT2 can be divided into <scene name='81/811717/Ntail_regions/1'>three different regions</scene>. Similar to 1BOB the N-terminal region of H4 is embedded in a cave between HAT1 and HAT2, though H4 mainly interacts with HAT1. The C-terminal helix of H4 is found inserted into LP2, the N-terminal helix, and C-terminal groove of HAT2. These interactions are strongly stabilized by salt bridge bonds between the histone and the complex. Previous studies suggest that H4K12 inserts into the active site of HAT1 to access AcCoA.<ref> | + | Once the complex has formed, histone 4 and AcetylCoA can begin to interact. The N-terminal segment of H4 that binds with HAT1/HAT2 can be divided into <scene name='81/811717/Ntail_regions/1'>three different regions</scene>. Similar to 1BOB the N-terminal region of H4 is embedded in a cave between HAT1 and HAT2, though H4 mainly interacts with HAT1. The C-terminal helix of H4 is found inserted into LP2, the N-terminal helix, and C-terminal groove of HAT2. These interactions are strongly stabilized by salt bridge bonds between the histone and the complex. Previous studies suggest that H4K12 inserts into the active site of HAT1 to access AcCoA.<ref name="Wu">PMID:22615379</ref> The 4PSW structure has H4K12 aligned with the active site and the AcCoA entering the concave groove from the opposite side. This allows the ε-amino group of H4K12 to contact the SH group of CoA. |
===Active Site=== | ===Active Site=== | ||
| - | The acetyl-CoA HAT1 active site is parallel to the C-terminal domain of the HAT1 protein. Acetyl-CoA fits structurally into the small binding site due to the kinked pantetheine group giving the molecule a bent confirmation. Once bound, most of the acetyl-CoA molecule is <scene name='81/811717/Tight_protein-ligand_intxn/1'>buried in the protein</scene> (~60%). Hydrophobic contacts, hydrogen bonds, and salt bridges help to stabilize the protein-ligand interaction. HAT1 protein-ligand contact is concentrated in three areas: C-terminal end of helix alpha 7, C terminal end of strand beta-14/loop Beta15-Alpha9, and N-terminal half of helix alpha 9 <ref | + | The acetyl-CoA HAT1 active site is parallel to the C-terminal domain of the HAT1 protein. Acetyl-CoA fits structurally into the small binding site due to the kinked pantetheine group giving the molecule a bent confirmation. Once bound, most of the acetyl-CoA molecule is <scene name='81/811717/Tight_protein-ligand_intxn/1'>buried in the protein</scene> (~60%). Hydrophobic contacts, hydrogen bonds, and salt bridges help to stabilize the protein-ligand interaction. HAT1 protein-ligand contact is concentrated in three areas: C-terminal end of helix alpha 7, C terminal end of strand beta-14/loop Beta15-Alpha9, and N-terminal half of helix alpha 9 <ref name=”Dutnall”>PMID:10384314</ref>. |
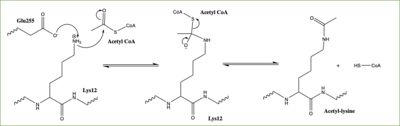
The β-methyl of the acetyl group interactions in the hydrophobic pocket formed by the side chain of residues: Ile-217,Pro-257, Phe-261. The carbonyl oxygen of the acetyl group hydrogen bonds with the aminde of the main chain Phe-220 and the sulfur of the acetyl-group interactions, as a hydrogen bond, with Asn-258. These interactions keep acetyl-CoA in the correct position of the active site for the transfer of the acetyl-group. | The β-methyl of the acetyl group interactions in the hydrophobic pocket formed by the side chain of residues: Ile-217,Pro-257, Phe-261. The carbonyl oxygen of the acetyl group hydrogen bonds with the aminde of the main chain Phe-220 and the sulfur of the acetyl-group interactions, as a hydrogen bond, with Asn-258. These interactions keep acetyl-CoA in the correct position of the active site for the transfer of the acetyl-group. | ||
Revision as of 16:23, 12 April 2019
Histone Acetyltransferase HAT1/HAT2 Complex, Saccharomyces cerevisiae
| |||||||||||