This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Asif Hossain/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 34: | Line 34: | ||
==Mechanism== | ==Mechanism== | ||
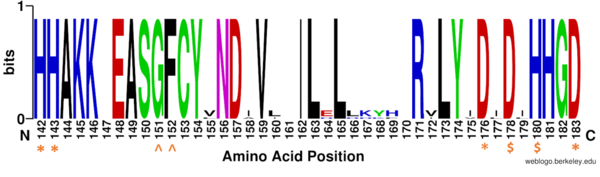
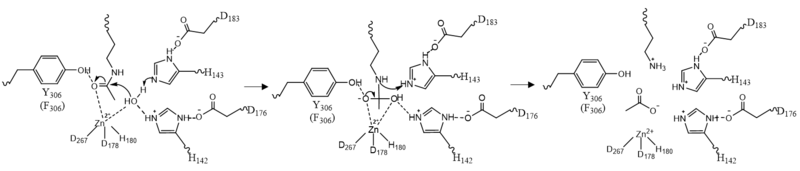
Once the substrate is bound to the binding pocket through interactions with <scene name='81/811087/Ligand_interaction/2'>Asp101, Phe152 and Phe208</scene>, the water molecule attacks the carbonyl carbon of the ε-amino-lysine sidechain of N-terminal core of histone proteins (Figure 2). This water molecule is recruited and stabilized by <scene name='81/811085/Dyads/2'>two catalytic dyads</scene>. The first dyad consists of His143 and Asp183. Asp183 interacts with His143 to shift electron density so that His143 may act as a general base to remove a proton from water. The second catalytic dyad consists of His142 and Asp176 and stabilizes the now deprotonated water molecule. A Zn<sup>2+</sup> ion also makes the water more acidic making it a better nucleophile. The tetrahedral intermediate is stabilized by the Zn<sup>2+</sup> ion as well as Tyr306. The amine group of the substrate's lysine acts as a general base and deprotonates His143. This drives the tetrahedral intermediate to collapse and expel the acetyl group to produce an acetate ion and a deacetylated lysine residue. <ref name="Seto, E., & Yoshida, M."> Seto, E., & Yoshida, M. (2014). Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harbor perspectives in biology, 6(4), a018713. https://doi.org/10.1101/cshperspect.a018713 </ref> | Once the substrate is bound to the binding pocket through interactions with <scene name='81/811087/Ligand_interaction/2'>Asp101, Phe152 and Phe208</scene>, the water molecule attacks the carbonyl carbon of the ε-amino-lysine sidechain of N-terminal core of histone proteins (Figure 2). This water molecule is recruited and stabilized by <scene name='81/811085/Dyads/2'>two catalytic dyads</scene>. The first dyad consists of His143 and Asp183. Asp183 interacts with His143 to shift electron density so that His143 may act as a general base to remove a proton from water. The second catalytic dyad consists of His142 and Asp176 and stabilizes the now deprotonated water molecule. A Zn<sup>2+</sup> ion also makes the water more acidic making it a better nucleophile. The tetrahedral intermediate is stabilized by the Zn<sup>2+</sup> ion as well as Tyr306. The amine group of the substrate's lysine acts as a general base and deprotonates His143. This drives the tetrahedral intermediate to collapse and expel the acetyl group to produce an acetate ion and a deacetylated lysine residue. <ref name="Seto, E., & Yoshida, M."> Seto, E., & Yoshida, M. (2014). Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harbor perspectives in biology, 6(4), a018713. https://doi.org/10.1101/cshperspect.a018713 </ref> | ||
| - | [[Image:Mech.PNG|800px||center||thumb|Figure 2: HDAC8 Mechanism:]] | + | [[Image:Mech.PNG|800px||center||thumb|Figure 2: HDAC8 Mechanism: Tyr306 was mutated to Phe306 to determine the crystal structure in the pdb file 2v5w.]] |
== Medical Relevance == | == Medical Relevance == | ||
Revision as of 20:40, 24 April 2019
Histone Deacetylase 8 (HDAC 8)
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 Vannini, A., Volpari, C., Gallinari, P., Jones, P., Mattu, M., Carfí, A., ... & Di Marco, S. (2007). Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8–substrate complex. EMBO reports, 8(9), 879-884. https://doi.org/10.1038/sj.embor.7401047
- ↑ DesJarlais, R., & Tummino, P. J. (2016). Role of histone-modifying enzymes and their complexes in regulation of chromatin biology. Biochemistry, 55(11), 1584-1599. https://doi.org/10.1021/acs.biochem.5b01210
- ↑ 3.0 3.1 3.2 3.3 3.4 Somoza J, Skene R. Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure, 12(7), 1325-1334.2004. https://doi.org/10.1016/j.str.2004.04.012
- ↑ Whitehead, L., Dobler, M. R., Radetich, B., Zhu, Y., Atadja, P. W., Claiborne, T., ... & Shao, W. (2011). Human HDAC isoform selectivity achieved via exploitation of the acetate release channel with structurally unique small molecule inhibitors. Bioorganic & medicinal chemistry, 19(15), 4626-4634. https://doi.org/10.1016/j.bmc.2011.06.030
- ↑ Seto, E., & Yoshida, M. (2014). Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harbor perspectives in biology, 6(4), a018713. https://doi.org/10.1101/cshperspect.a018713
- ↑ Eckschlager T, Plch, J, Stiborova M, Hrabeta J.Histone deacetylase inhibitors as anticancer drugs. International journal of molecular sciences, 18(7), 1414. 2017. https://dx.doi.org/10.3390%2Fijms18071414
- ↑ Vannini, A., Volpari, C., Filocamo, G., Casavola, E. C., Brunetti, M., Renzoni, D., ... & Steinkühler, C. (2004). Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proceedings of the National Academy of Sciences, 101(42), 15064-15069. https://dx.doi.org/10.1073%2Fpnas.0404603101