This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Madeleine Wilson/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 16: | Line 16: | ||
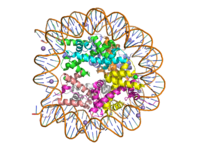
The secondary structure is composed of 10% <scene name='81/811086/Helices/3'>helices</scene> and 37% <scene name='81/811086/Beta_sheets/3'>beta sheets</scene>. The helical composition includes 3 <scene name='81/811086/Alpha_helices/4'>alpha helices</scene>, with two residing in the SET domain and one in the C-terminal domain. The alpha helices in the SET domain are two turns while the C-terminal helix is by far the largest with 4 turns. There are also 2 <scene name='81/811086/3-10_helices/4'>3-10 helices</scene> in the SET domain which are each one turn. There are 21 total beta strands which reside in the N-terminal domain and the SET domain. The beta strands are primarily anti-parallel and multiple antiparallel strands are connected by <scene name='81/811086/Type_i_beta_turns/3'>Type I</scene> and <scene name='81/811086/Type_ii_beta_turns/2'>Type II</scene> beta turns. | The secondary structure is composed of 10% <scene name='81/811086/Helices/3'>helices</scene> and 37% <scene name='81/811086/Beta_sheets/3'>beta sheets</scene>. The helical composition includes 3 <scene name='81/811086/Alpha_helices/4'>alpha helices</scene>, with two residing in the SET domain and one in the C-terminal domain. The alpha helices in the SET domain are two turns while the C-terminal helix is by far the largest with 4 turns. There are also 2 <scene name='81/811086/3-10_helices/4'>3-10 helices</scene> in the SET domain which are each one turn. There are 21 total beta strands which reside in the N-terminal domain and the SET domain. The beta strands are primarily anti-parallel and multiple antiparallel strands are connected by <scene name='81/811086/Type_i_beta_turns/3'>Type I</scene> and <scene name='81/811086/Type_ii_beta_turns/2'>Type II</scene> beta turns. | ||
| - | |||
| - | ===The N-terminal Domain=== | ||
| - | [[Image:Clustal.png|200px|right|thumb|Figure 1. Clustal alignment of N-terminals]] | ||
| - | The N-terminal domain has no notable function in regard to the activity of KMT; in addition to this, no recent studies have been conducted to discover what the exact function of this region is. As this is a highly conserved region across species, it may be hypothesized this domain plays an integral part in stability. | ||
===The Active Site=== | ===The Active Site=== | ||
| Line 31: | Line 27: | ||
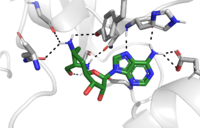
The C-terminal domain of lysine methyltransferase is very important for the catalytic activity of the enzyme. The structures of the <scene name='81/811091/C_terminal_domain/1'>C-terminal domain (residues 345-366)</scene> serve the role of stabilizing the structures in the <scene name='81/811091/C_terminal_domain/16'>SET7 domain (residues 193-344)</scene> in the correct orientation for a reaction in the active site.<ref name="Xiao" /> Hydrophobic interactions in the C-terminal domain are mainly responsible for stabilizing the access channel for the lysine methylation site on histone H3. Residues 337-349 create a beta-hairpin structure that stabilizes the orientation of two tyrosine residues, Tyr335 and Tyr337, that form the opening to the lysine access channel. Furthermore, the hydrophobic packing of alpha-helix 3 against beta-sheet 19, specifically <scene name='81/811091/C_terminal_domain/13'>residues Leu357 and Phe 299</scene>, stabilize the orientation of the <scene name='81/811091/C_terminal_domain/14'>SAM cofactor</scene> so that its methyl donating group is oriented toward the lysine access channel. The orientation of the SAM cofactor is further stabilized with its hydrophobic interactions with C-terminal domain residues <scene name='81/811091/C_terminal_domain/18'>Trp352 and Glu356</scene>. <ref name="Xiao" /> | The C-terminal domain of lysine methyltransferase is very important for the catalytic activity of the enzyme. The structures of the <scene name='81/811091/C_terminal_domain/1'>C-terminal domain (residues 345-366)</scene> serve the role of stabilizing the structures in the <scene name='81/811091/C_terminal_domain/16'>SET7 domain (residues 193-344)</scene> in the correct orientation for a reaction in the active site.<ref name="Xiao" /> Hydrophobic interactions in the C-terminal domain are mainly responsible for stabilizing the access channel for the lysine methylation site on histone H3. Residues 337-349 create a beta-hairpin structure that stabilizes the orientation of two tyrosine residues, Tyr335 and Tyr337, that form the opening to the lysine access channel. Furthermore, the hydrophobic packing of alpha-helix 3 against beta-sheet 19, specifically <scene name='81/811091/C_terminal_domain/13'>residues Leu357 and Phe 299</scene>, stabilize the orientation of the <scene name='81/811091/C_terminal_domain/14'>SAM cofactor</scene> so that its methyl donating group is oriented toward the lysine access channel. The orientation of the SAM cofactor is further stabilized with its hydrophobic interactions with C-terminal domain residues <scene name='81/811091/C_terminal_domain/18'>Trp352 and Glu356</scene>. <ref name="Xiao" /> | ||
| + | |||
| + | ===The N-terminal Domain=== | ||
| + | [[Image:Clustal.png|200px|right|thumb|Figure 1. Clustal alignment of N-terminals]] | ||
| + | The N-terminal domain has no notable function in regard to the activity of KMT; in addition to this, no recent studies have been conducted to discover what the exact function of this region is. As this is a highly conserved region across species, it may be hypothesized this domain plays an integral part in stability. | ||
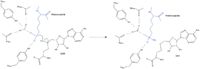
==Inhibitors== | ==Inhibitors== | ||
Revision as of 00:08, 26 April 2019
Histone Lysine Methyltransferase: Gene Activator
| |||||||||||
References
- ↑ DesJarlais R, Tummino PJ. Role of Histone-Modifying Enzymes and Their Complexes in Regulation of Chromatin Biology. Biochemistry. 2016 Mar 22;55(11):1584-99. doi: 10.1021/acs.biochem.5b01210. Epub , 2016 Jan 26. PMID:26745824 doi:http://dx.doi.org/10.1021/acs.biochem.5b01210
- ↑ 2.0 2.1 doi: https://dx.doi.org/10.1016/j.apsb.2013.04.007
- ↑ 3.0 3.1 Dong X, Weng Z. The correlation between histone modifications and gene expression. Epigenomics. 2013 Apr;5(2):113-6. doi: 10.2217/epi.13.13. PMID:23566087 doi:http://dx.doi.org/10.2217/epi.13.13
- ↑ 4.0 4.1 Del Rizzo PA, Trievel RC. Substrate and product specificities of SET domain methyltransferases. Epigenetics. 2011 Sep 1;6(9):1059-67. doi: 10.4161/epi.6.9.16069. Epub 2011 Sep, 1. PMID:21847010 doi:http://dx.doi.org/10.4161/epi.6.9.16069
- ↑ 5.0 5.1 5.2 5.3 5.4 5.5 Xiao B, Jing C, Wilson JR, Walker PA, Vasisht N, Kelly G, Howell S, Taylor IA, Blackburn GM, Gamblin SJ. Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature. 2003 Feb 6;421(6923):652-6. Epub 2003 Jan 22. PMID:12540855 doi:10.1038/nature01378
- ↑ Schluckebier G, Kozak M, Bleimling N, Weinhold E, Saenger W. Differential binding of S-adenosylmethionine S-adenosylhomocysteine and Sinefungin to the adenine-specific DNA methyltransferase M.TaqI. J Mol Biol. 1997 Jan 10;265(1):56-67. PMID:8995524 doi:http://dx.doi.org/10.1006/jmbi.1996.0711
- ↑ Tamura R, Doi S, Nakashima A, Sasaki K, Maeda K, Ueno T, Masaki T. Inhibition of the H3K4 methyltransferase SET7/9 ameliorates peritoneal fibrosis. PLoS One. 2018 May 3;13(5):e0196844. doi: 10.1371/journal.pone.0196844., eCollection 2018. PMID:29723250 doi:http://dx.doi.org/10.1371/journal.pone.0196844
Student Contributors
Lauryn Padgett, Alexandra Pentala, Madeleine Wilson