Introduction
Histone deacetylase 8 (HDAC8) is an enzyme that plays a role in controlling gene expression. Specifically, HDAC8 removes an acetyl group off of the ε-amino-Lys 382 of Histone 4's N-terminal core.[1] Histones consist of eight monomers to form an octomer complex. Each histone has a positive charge which allows interaction with negatively-charged DNA. This prevents transcription factors from accessing DNA, thus, decreasing gene expression. Chromatin remodeling by the addition or removal of an acetyl group on a histone is an example of epigenetic regulation. Histone Acetylase 1 (HAT1) catalyzes the addition of an acetyl group onto a histone. The lack of charge on the acetyl group weakens the interaction between DNA and histones which allows transcription factors to access the DNA to increase gene expression. HDAC8 reverses this reaction by catalyzing the removal of these acetyl groups from the Lys to reclaim the positive charge of the histone. This allows the histone to interact with the negative charge on the DNA. As a result, DNA binds more tightly to the histone protein, repressing transcription and gene expression.
HDAC Enzymes and Homology
There are four major classes of HDAC proteins (I,II, III, and IV). Other than the Class III “sirtuins” that utilize a NAD+ cofactor-dependent mechanism, all other HDAC classes use Zn2+-assisted catalysis through mechanisms (Figure 3) reminiscent of a typical serine protease.[2] While Classes I, II, and IV do have some major distinctions such as size of the protein, in general, they share homology at the catalytic site. HDAC 8 is classified as a Class I HDAC alongside HDACs 1-3. In fact, within Class I HDACs, there are many invariant residues involved in the catalytic site (such as His-Asp dyads), Zn-binding, and ligand binding pocket (such as Asp101) (Figure 1). [1]
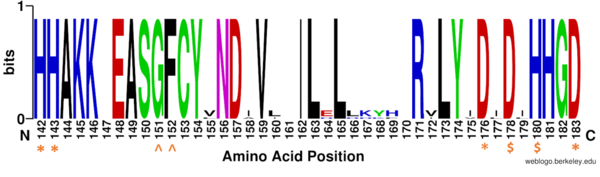
Figure 1: Weblogo representation comparing conservation of residues (143-182 in HDAC8) to homologous sequences in all class I HDACs. Nearly all active site residues (asterisk), zinc binding (dollar), and binding pocket residues (caret) are conserved across all class I HDACs. Other conserved residues not shown include active site residue Tyr306, zinc binding residue Asp267, and binding pocket residue Asp101. Nonconserved residues from 158 to l70 are part of an α-helix that moves outward from the active side before looping back around to the active site.
HDAC8 Structure
The crystal structure of human HDAC8 was determined using x-ray crystallography at a 2.0Å resolution. [1] The structure includes two structural K ion and one catalytic Zn ion. HDAC8 is bound to a p-53 as opposed to the natural histone substrate. This peptide includes a likely used in past kinetic assays. The HDAC8 is made up of a with eight parallel β-strands located between 13 . The HDAC8 consists of 377 amino acids. [3]
Zinc Ion
The pentacoordinated Zn2+ ion involved in the metalloenzyme catalysis is tethered to the protein through interactions with . This positions the metal ion to favorably interact with the catalytic water and acetylated lysine substrate. [1] The Zn2+ ion lowers the pKa of a water molecule that activates it as a nucleophile. By binding to both the nucleophile and the substrate simultaneously, the Zn2+ ion also assists the deacetylation process by lowering the entropy of the reaction. This polarizes the carbonyl of the acetyl-lysine and stabilizes the transition state.[3]
Key Residues
The of HDAC8 is composed of 2 catalytic dyads: , which activate the catalytic water nucleophile. A Tyr306, through mutation to Phe in the pdb file 2v5w (modeled in the overall view) was observed to render the protein mostly inactive. Thus, it has been hypothesized that the this residue is critical for stabilization of the transition state with the Zn2+ ion. This mutation allowed the determination of the crystal structure of HDAC8 interacting the ligand. [1]
Binding Pocket
By encasing the nonpolar, four-carbon side-chain of the Lys residue on the ligand, Phe152 and Phe208 engage in hydrophobic Van der Waals interactions with the ligand at different ends of the . Trp141 and Met274 contribute to the overall shape through general hydrophobic interactions.[4] Finally, the carbonyl oxygen of hydrogen bonds with the amide hydrogen of the acetylated lysine to further interact with the ligand in the relatively hydrophobic tunnel.[1]
At the rim of the active site, is involved in two hydrogen bonds between its own carbonyl oxygens and two consecutive amide hydrogens of incoming peptide derived ligand. This forces the ligand to assume a cis-conformation. In addition, extensive interactions among many other polar atoms near the rim of the active site help keep the ligand lodged in the hydrophobic tunnel.[1]
Additional Features
There are two potassium ions bound in the HDAC8 structure. Potassium 1 is 7Å away from the active site while potassium 2 lies toward the exterior of the HDAC8.[5]. It is suggested that potassium 1 is of interest to the active site of HDAC8 because it is tethered to the enzyme by the main chain carbonyl oxygens of Asp178 and His180 which stabilizes the Zn2+ in the active site. Furthermore, the potassium ion increases the positive charge in the active site and this could help stabilize the oxyanion hole that is formed in the transition state.[5]
(amino acid residues 30-36) makes up a significant part of the active site pocket. It is suggested that this loop has high flexibility that enables HDAC8 to more efficiently adjust binding to different ligands. [3]
The activity of HDAC8 is regulated by phosphorylation at by protein kinase A (PKA) as the phosphorylation has been shown to decrease the activity of HDAC8.[3] It is suggested that the phosphorylation of Ser39 can interfere with the interaction between HDAC8 and the target histone because Ser39 is at the periphery of HDAC8, approximately 20Å from the active site. The interaction is disrupted because the phosphorylated Ser39 elicits a structural rearrangement that reaches the active site and disrupts enzyme activity. Ser39 elicits the rearrangement by interactions with structural elements in the conformational active loop L1, such as Lys36. Therefore, the Ser39 phosphorylation by PKA is likely inducing a conformational change of the L1 loop that prohibits a substrate binding.[3]
Mechanism
Once the substrate is bound to the binding pocket through interactions with , the water molecule attacks the carbonyl carbon of the ε-amino-lysine sidechain of N-terminal core of histone proteins (Figure 2). This water molecule is recruited and stabilized by . The first dyad consists of His143 and Asp183. Asp183 interacts with His143 to shift electron density so that His143 may act as a general base to remove a proton from water. The second catalytic dyad consists of His142 and Asp176 and stabilizes the now deprotonated water molecule. A Zn2+ ion also makes the water more acidic making it a better nucleophile. The tetrahedral intermediate is stabilized by the Zn2+ ion as well as Tyr306. The amine group of the substrate's lysine acts as a general base and deprotonates His143. This drives the tetrahedral intermediate to collapse and expel the acetyl group to produce an acetate ion and a deacetylated lysine residue. [6]
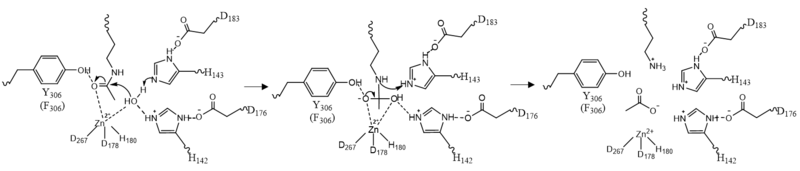
Figure 2: HDAC8 Mechanism: Tyr306 was mutated to Phe306 to determine the crystal structure in the pdb file 2v5w.
Medical Relevance
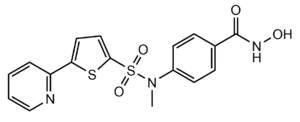
Figure 3: Structure of "Compound 1," a Hydroxamic Inhibitor
Besides controlling gene regulation through deacetylation of histones, HDAC8 also regulates other non-histone proteins such as chaperones, hormone receptors and signaling molecules. [7] Thus, it has influences on protein stability and interactions between other proteins and DNA. HDAC8 can therefore affect the regulation of cell proliferation and cell death. These processes are typically being altered in cancer cells and that makes HDAC enzymes an interesting potential target for cancer drugs. HDAC inhibitors have been shown to be promising cancer drug agent as the HDAC inhibitors (HDACi) cease tumor growth in cancer cells by either making them differentiate, undergo apoptosis or upregulate cell cycle arrest proteins. [7] One way the HDAC inhibitors cease tumor growth is by the reactivation of the transcription factor, RUNX3, a known tumor suppressor. HDACi increases the acetylation of the protein and as the stability of RUNX3 is dependent on the acetylation status of the protein, the increased acetylation or HDAC inhibition will enhance the protein stability. A number of HDAC inhibitors have been purified from natural sources or synthesized and at least four structurally different inhibitor classes have been characterized: hydroxamates, cyclic peptides, aliphatic acids and benzamides. The Vorinostat (within the hydroxamate class) has been FDA-approved for treatment of cancer. A HDAC class 1 hydroxamic acid, compound 1 (Figure 3), in a bidendate fashion while making hydrogen bonds to important residues as His142, His143 and Tyr306 at the active site of HDAC8. Thus, the HDAC inhibitors can be used as antagonists to prevent the functioning of HDAC8 in cancer treatment. [5]



