User:Asif Hossain/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 19: | Line 19: | ||
===Key Residues=== | ===Key Residues=== | ||
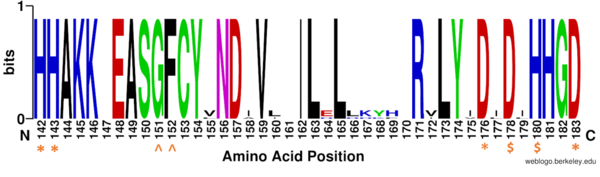
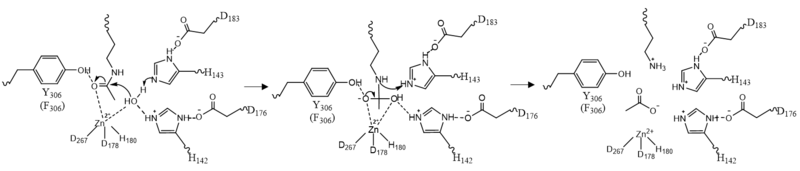
| - | The <scene name='81/811085/Active_site/ | + | The <scene name='81/811085/Active_site/13'>active site</scene> of HDAC8 is composed of 2 catalytic dyads: <scene name='81/811085/Dyads/5'>His143/Asp183 and His142/Asp176</scene>, which activate the catalytic water nucleophile. A Tyr306, through mutation to Phe in the pdb file 2v5w (modeled in the overall view) was observed to render the protein mostly inactive. Thus, it has been hypothesized that the this residue is critical for stabilization of the transition state with the Zn<sup>2+</sup> ion. This mutation allowed the determination of the crystal structure of HDAC8 interacting the ligand. <ref name="Vannini, A., Volpari, C., Gallinari, P.">Vannini, A., Volpari, C., Gallinari, P., Jones, P., Mattu, M., Carfí, A., ... & Di Marco, S. (2007). Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8–substrate complex. EMBO reports, 8(9), 879-884. https://doi.org/10.1038/sj.embor.7401047 </ref> |
===Binding Pocket=== | ===Binding Pocket=== | ||
| Line 29: | Line 29: | ||
There are two potassium ions bound in the HDAC8 structure. Potassium 1 is 7Å away from the active site while potassium 2 lies toward the exterior of the HDAC8.<ref name="Vannini, A., Volpari, C., Filocamo, G.">Vannini, A., Volpari, C., Filocamo, G., Casavola, E. C., Brunetti, M., Renzoni, D., ... & Steinkühler, C. (2004). Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proceedings of the National Academy of Sciences, 101(42), 15064-15069. https://dx.doi.org/10.1073%2Fpnas.0404603101</ref>. It is suggested that potassium 1 is of interest to the active site of HDAC8 because it is tethered to the enzyme by the main chain carbonyl oxygens of Asp178 and His180 which stabilizes the Zn<sup>2+</sup> in the active site. Furthermore, the potassium ion increases the positive charge in the active site and this could help stabilize the oxyanion hole that is formed in the transition state.<ref name="Vannini, A., Volpari, C., Filocamo, G.">Vannini, A., Volpari, C., Filocamo, G., Casavola, E. C., Brunetti, M., Renzoni, D., ... & Steinkühler, C. (2004). Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proceedings of the National Academy of Sciences, 101(42), 15064-15069. https://dx.doi.org/10.1073%2Fpnas.0404603101</ref> | There are two potassium ions bound in the HDAC8 structure. Potassium 1 is 7Å away from the active site while potassium 2 lies toward the exterior of the HDAC8.<ref name="Vannini, A., Volpari, C., Filocamo, G.">Vannini, A., Volpari, C., Filocamo, G., Casavola, E. C., Brunetti, M., Renzoni, D., ... & Steinkühler, C. (2004). Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proceedings of the National Academy of Sciences, 101(42), 15064-15069. https://dx.doi.org/10.1073%2Fpnas.0404603101</ref>. It is suggested that potassium 1 is of interest to the active site of HDAC8 because it is tethered to the enzyme by the main chain carbonyl oxygens of Asp178 and His180 which stabilizes the Zn<sup>2+</sup> in the active site. Furthermore, the potassium ion increases the positive charge in the active site and this could help stabilize the oxyanion hole that is formed in the transition state.<ref name="Vannini, A., Volpari, C., Filocamo, G.">Vannini, A., Volpari, C., Filocamo, G., Casavola, E. C., Brunetti, M., Renzoni, D., ... & Steinkühler, C. (2004). Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proceedings of the National Academy of Sciences, 101(42), 15064-15069. https://dx.doi.org/10.1073%2Fpnas.0404603101</ref> | ||
| - | <scene name='81/811087/Active_site_loop_1_s30-k36/ | + | <scene name='81/811087/Active_site_loop_1_s30-k36/14'>N-Terminus L1 loop</scene>(amino acid residues 30-36) makes up a significant part of the active site pocket. It is suggested that this loop has high flexibility that enables HDAC8 to more efficiently adjust binding to different ligands. <ref name="Somoza">Somoza J, Skene R. Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure, 12(7), 1325-1334.2004. https://doi.org/10.1016/j.str.2004.04.012 </ref> |
The activity of HDAC8 is regulated by phosphorylation at <scene name='81/811087/Active_site_loop_1_s30-k36/13'>Ser39</scene> by [https://en.wikipedia.org/wiki/Protein_kinase_A protein kinase A (PKA)] as the phosphorylation has been shown to decrease the activity of HDAC8.<ref name="Somoza">Somoza J, Skene R. Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure, 12(7), 1325-1334.2004. https://doi.org/10.1016/j.str.2004.04.012 </ref> It is suggested that the phosphorylation of Ser39 can interfere with the interaction between HDAC8 and the target histone because Ser39 is at the periphery of HDAC8, approximately 20Å from the active site. The interaction is disrupted because the phosphorylated Ser39 elicits a structural rearrangement that reaches the active site and disrupts enzyme activity. Ser39 elicits the rearrangement by interactions with structural elements in the conformational active loop L1, such as Lys36. Therefore, the Ser39 phosphorylation by PKA is likely inducing a conformational change of the L1 loop that prohibits a substrate binding.<ref name="Somoza">Somoza J, Skene R. Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure, 12(7), 1325-1334.2004. https://doi.org/10.1016/j.str.2004.04.012 </ref> | The activity of HDAC8 is regulated by phosphorylation at <scene name='81/811087/Active_site_loop_1_s30-k36/13'>Ser39</scene> by [https://en.wikipedia.org/wiki/Protein_kinase_A protein kinase A (PKA)] as the phosphorylation has been shown to decrease the activity of HDAC8.<ref name="Somoza">Somoza J, Skene R. Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure, 12(7), 1325-1334.2004. https://doi.org/10.1016/j.str.2004.04.012 </ref> It is suggested that the phosphorylation of Ser39 can interfere with the interaction between HDAC8 and the target histone because Ser39 is at the periphery of HDAC8, approximately 20Å from the active site. The interaction is disrupted because the phosphorylated Ser39 elicits a structural rearrangement that reaches the active site and disrupts enzyme activity. Ser39 elicits the rearrangement by interactions with structural elements in the conformational active loop L1, such as Lys36. Therefore, the Ser39 phosphorylation by PKA is likely inducing a conformational change of the L1 loop that prohibits a substrate binding.<ref name="Somoza">Somoza J, Skene R. Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure, 12(7), 1325-1334.2004. https://doi.org/10.1016/j.str.2004.04.012 </ref> | ||
Revision as of 17:10, 26 April 2019
Histone Deacetylase 8 (HDAC 8)
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 Vannini, A., Volpari, C., Gallinari, P., Jones, P., Mattu, M., Carfí, A., ... & Di Marco, S. (2007). Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8–substrate complex. EMBO reports, 8(9), 879-884. https://doi.org/10.1038/sj.embor.7401047
- ↑ DesJarlais, R., & Tummino, P. J. (2016). Role of histone-modifying enzymes and their complexes in regulation of chromatin biology. Biochemistry, 55(11), 1584-1599. https://doi.org/10.1021/acs.biochem.5b01210
- ↑ 3.0 3.1 3.2 3.3 3.4 Somoza J, Skene R. Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure, 12(7), 1325-1334.2004. https://doi.org/10.1016/j.str.2004.04.012
- ↑ Whitehead, L., Dobler, M. R., Radetich, B., Zhu, Y., Atadja, P. W., Claiborne, T., ... & Shao, W. (2011). Human HDAC isoform selectivity achieved via exploitation of the acetate release channel with structurally unique small molecule inhibitors. Bioorganic & medicinal chemistry, 19(15), 4626-4634. https://doi.org/10.1016/j.bmc.2011.06.030
- ↑ 5.0 5.1 5.2 Vannini, A., Volpari, C., Filocamo, G., Casavola, E. C., Brunetti, M., Renzoni, D., ... & Steinkühler, C. (2004). Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proceedings of the National Academy of Sciences, 101(42), 15064-15069. https://dx.doi.org/10.1073%2Fpnas.0404603101
- ↑ Seto, E., & Yoshida, M. (2014). Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harbor perspectives in biology, 6(4), a018713. https://doi.org/10.1101/cshperspect.a018713
- ↑ 7.0 7.1 Eckschlager T, Plch, J, Stiborova M, Hrabeta J.Histone deacetylase inhibitors as anticancer drugs. International journal of molecular sciences, 18(7), 1414. 2017. https://dx.doi.org/10.3390%2Fijms18071414