This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2vgb
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
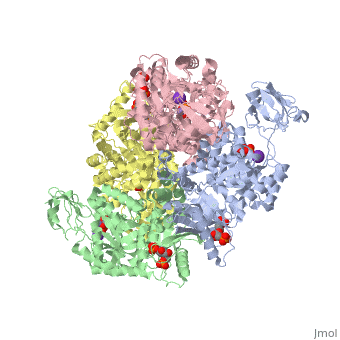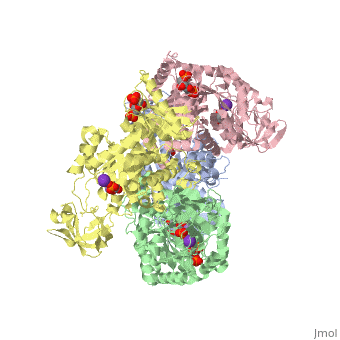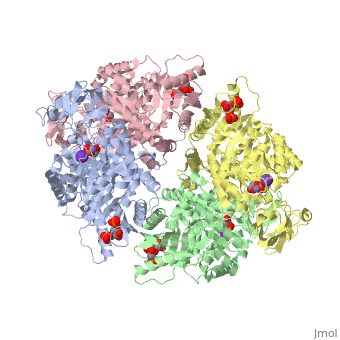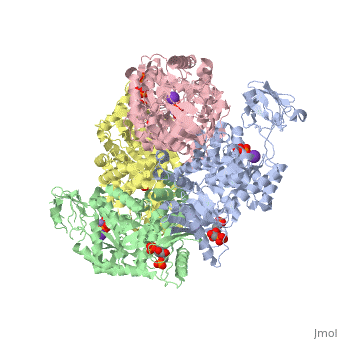
<StructureSection load='2vgb' size='340' side='right'caption='[[2vgb]], [[Resolution|resolution]] 2.73Å' scene=''> | <StructureSection load='2vgb' size='340' side='right'caption='[[2vgb]], [[Resolution|resolution]] 2.73Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[2vgb]] is a 4 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[2vgb]] is a 4 chain structure with sequence from [https://en.wikipedia.org/wiki/Human Human]. This structure supersedes the now removed PDB entry [http://oca.weizmann.ac.il/oca-bin/send-pdb?obs=1&id=1liu 1liu]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2VGB OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2VGB FirstGlance]. <br> |
| - | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=FBP:BETA-FRUCTOSE-1,6-DIPHOSPHATE'>FBP</scene>, <scene name='pdbligand=K:POTASSIUM+ION'>K</scene>, <scene name='pdbligand=MN:MANGANESE+(II)+ION'>MN</scene>, <scene name='pdbligand=PGA:2-PHOSPHOGLYCOLIC+ACID'>PGA</scene></td></tr> | + | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=FBP:BETA-FRUCTOSE-1,6-DIPHOSPHATE'>FBP</scene>, <scene name='pdbligand=K:POTASSIUM+ION'>K</scene>, <scene name='pdbligand=MN:MANGANESE+(II)+ION'>MN</scene>, <scene name='pdbligand=PGA:2-PHOSPHOGLYCOLIC+ACID'>PGA</scene></td></tr> |
| - | <tr id='related'><td class="sblockLbl"><b>[[Related_structure|Related:]]</b></td><td class="sblockDat">[[1liy|1liy]], [[1liu|1liu]], [[1liw|1liw]], [[1lix|1lix]]</td></tr> | + | <tr id='related'><td class="sblockLbl"><b>[[Related_structure|Related:]]</b></td><td class="sblockDat"><div style='overflow: auto; max-height: 3em;'>[[1liy|1liy]], [[1liu|1liu]], [[1liw|1liw]], [[1lix|1lix]]</div></td></tr> |
| - | <tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[https://en.wikipedia.org/wiki/Pyruvate_kinase Pyruvate kinase], with EC number [https://www.brenda-enzymes.info/php/result_flat.php4?ecno=2.7.1.40 2.7.1.40] </span></td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2vgb FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2vgb OCA], [https://pdbe.org/2vgb PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2vgb RCSB], [https://www.ebi.ac.uk/pdbsum/2vgb PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2vgb ProSAT]</span></td></tr> |
</table> | </table> | ||
== Disease == | == Disease == | ||
| - | [[ | + | [[https://www.uniprot.org/uniprot/KPYR_HUMAN KPYR_HUMAN]] Defects in PKLR are the cause of pyruvate kinase hyperactivity (PKHYP) [MIM:[https://omim.org/entry/102900 102900]]; also known as high red cell ATP syndrome. This autosomal dominant phenotype is characterized by increase of red blood cell ATP.<ref>PMID:9090535</ref> Defects in PKLR are the cause of pyruvate kinase deficiency of red cells (PKRD) [MIM:[https://omim.org/entry/266200 266200]]. A frequent cause of hereditary non-spherocytic hemolytic anemia. Clinically, pyruvate kinase-deficient patients suffer from a highly variable degree of chronic hemolysis, ranging from severe neonatal jaundice and fatal anemia at birth, severe transfusion-dependent chronic hemolysis, moderate hemolysis with exacerbation during infection, to a fully compensated hemolysis without apparent anemia. |
== Function == | == Function == | ||
| - | [[ | + | [[https://www.uniprot.org/uniprot/KPYR_HUMAN KPYR_HUMAN]] Plays a key role in glycolysis (By similarity). |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 35: | Line 35: | ||
==See Also== | ==See Also== | ||
*[[Pyruvate Kinase|Pyruvate Kinase]] | *[[Pyruvate Kinase|Pyruvate Kinase]] | ||
| + | *[[Pyruvate kinase 3D structures|Pyruvate kinase 3D structures]] | ||
== References == | == References == | ||
<references/> | <references/> | ||
Revision as of 10:45, 7 July 2021
HUMAN ERYTHROCYTE PYRUVATE KINASE
| |||||||||||
Categories: Human | Large Structures | Pyruvate kinase | Abraham, D J | Bianchi, P | Chiarelli, L | Dolzan, M | Fortin, R | Galizzi, A | Mattevi, A | Valentini, G | Wang, C | Zanella, A | Disease mutation | Glycolysis | Kinase | Magnesium | Metal-binding | Phosphorylation | Pyruvate | Pyruvate kinase in the active r-state | Transferase