This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Hyaluronidase
From Proteopedia
(Difference between revisions)
| Line 12: | Line 12: | ||
== Structural highlights == | == Structural highlights == | ||
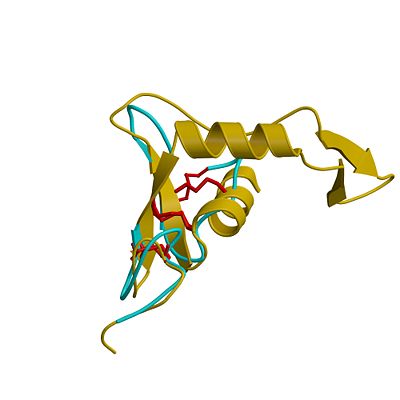
| - | HU structure contains an <scene name='49/497055/Cv/ | + | HU structure contains an <scene name='49/497055/Cv/8'>N-terminal, linker and C-terminal domains</scene>. The active site is in a cleft in the N-terminal domain. Two HUA molecules are bound in the active site making contacts with some HU residues and multiple contacts with water molecules<ref>PMID:10843845</ref>. |
| - | <scene name='49/497055/Cv/ | + | <scene name='49/497055/Cv/9'>1st active site of HU</scene>. Water molecules are shown as red spheres. |
| - | <scene name='49/497055/Cv/ | + | <scene name='49/497055/Cv/10'>2nd active site of HU</scene>. |
| - | <scene name='49/497055/Cv/ | + | <scene name='49/497055/Cv/11'>Dimethylarsinate binding site</scene>. |
</StructureSection> | </StructureSection> | ||
Revision as of 12:57, 19 May 2019
| |||||||||||
3D structures of hyaluronidase
Updated on 19-May-2019
Reference
- ↑ Chao KL, Muthukumar L, Herzberg O. Structure of human hyaluronidase-1, a hyaluronan hydrolyzing enzyme involved in tumor growth and angiogenesis. Biochemistry. 2007 Jun 12;46(23):6911-20. Epub 2007 May 16. PMID:17503783 doi:10.1021/bi700382g
- ↑ Ponnuraj K, Jedrzejas MJ. Mechanism of hyaluronan binding and degradation: structure of Streptococcus pneumoniae hyaluronate lyase in complex with hyaluronic acid disaccharide at 1.7 A resolution. J Mol Biol. 2000 Jun 16;299(4):885-95. PMID:10843845 doi:10.1006/jmbi.2000.3817
Created with the participation of Osnat Herzberg, Eran Hodis, Joel L. Sussman, Jaime Prilusky.