This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
LdtMt2
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
Expression System: Escherichia coli BL21(DE3) | Expression System: Escherichia coli BL21(DE3) | ||
' scene='81/817533/All/1'> | ' scene='81/817533/All/1'> | ||
| - | + | ||
The L,D-transpeptidase <scene name='81/817533/All/1'>LdtMt2</scene> is an enzyme that catalyzes the formation of peptidoglycan crosslinking in ''Mycobaterium tuberculosis''. | The L,D-transpeptidase <scene name='81/817533/All/1'>LdtMt2</scene> is an enzyme that catalyzes the formation of peptidoglycan crosslinking in ''Mycobaterium tuberculosis''. | ||
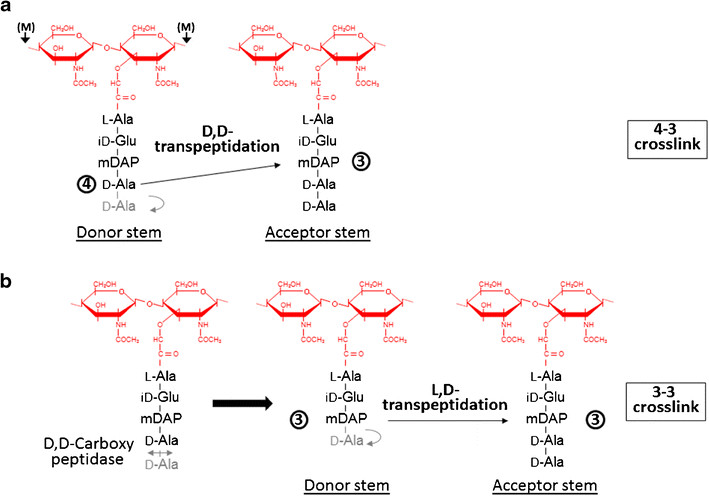
| - | Formation of the most common type of crosslink in peptidoglycan, the (D,D) 4->3 linkage, is catalyzed by the [D,d-transpeptidase]. These enzymes generate 4 -> 3 transpeptidase linkages between the fourth amino acid (D-alanine) of one chain and the third amino acid (meso-diaminopimelic acid) of an adjacent chain. A second type of crosslink, the (L,D) 3->3 linkage is catalysed by L,D-transpeptidases such as Mtb L,D-transpeptidase LdtMt2 of ''Mycobaterium tuberculosis''. These enzymes transfer the peptide bond between the third residue (meso-diaminopimelic acid) of a tetrapeptide donor stem to the side-chain amide group of the third residue (meso-diaminopimelic acid) of an adjacent acceptor stem. In both types of transpeptidases, the catalysis proceeds by a two-step mechanism: acylation of the enzyme by the penultimate peptide of the donor stem with the release of the stem C-terminal residue, followed by deacylation of this acyl-enzyme intermediate by an acceptor stem. | + | Formation of the most common type of crosslink in peptidoglycan, the (D,D) 4->3 linkage, is catalyzed by the [[D,d-transpeptidase]]. These enzymes generate 4 -> 3 transpeptidase linkages between the fourth amino acid (D-alanine) of one chain and the third amino acid (meso-diaminopimelic acid) of an adjacent chain. A second type of crosslink, the (L,D) 3->3 linkage is catalysed by L,D-transpeptidases such as Mtb L,D-transpeptidase LdtMt2 of ''Mycobaterium tuberculosis''. These enzymes transfer the peptide bond between the third residue (meso-diaminopimelic acid) of a tetrapeptide donor stem to the side-chain amide group of the third residue (meso-diaminopimelic acid) of an adjacent acceptor stem. In both types of transpeptidases, the catalysis proceeds by a two-step mechanism: acylation of the enzyme by the penultimate peptide of the donor stem with the release of the stem C-terminal residue, followed by deacylation of this acyl-enzyme intermediate by an acceptor stem. |
[[Image:Figure transpeptidases.png]] | [[Image:Figure transpeptidases.png]] | ||
| Line 21: | Line 21: | ||
== Structure == | == Structure == | ||
| - | The LdtMt2 precursor consists of 408 amino acid residues that form the transmembrane domain (Met1-Ala34) and the chain of the enzyme (Cys35-Ala408), which can be divided into three domains: two non-catalytic Ig-like <scene name='81/817533/Domaina/1'>Domain A</scene>, <scene name='81/817533/Domainb/1'>Domain B</scene> (residues Ala55-Ser147 and Pro148- Gly250, respectively) and the <scene name='81/817533/Domaincd/1'>catalytic domain</scene> (residues Asp251-Ala408) with transpeptidase activity. Domain A e Domain B | + | The LdtMt2 precursor consists of 408 amino acid residues that form the transmembrane domain (Met1-Ala34) and the chain of the enzyme (Cys35-Ala408), which can be divided into <scene name='81/817533/Second/3'>three domains</scene>: two non-catalytic Ig-like <scene name='81/817533/Domaina/1'>Domain A</scene>, <scene name='81/817533/Domainb/1'>Domain B</scene> (residues Ala55-Ser147 and Pro148- Gly250, respectively) and the <scene name='81/817533/Domaincd/1'>catalytic domain</scene> (residues Asp251-Ala408) with transpeptidase activity. Domain A e Domain B displays a variant to the immunoglobulin fold built up by a sandwich of two antiparallel sheets. And the catalytic domain consists of a beta-sandwich with two mixed beta sheets ErfK/YbiS/YhnG fold. |
| - | A short linker (residues 251–253) joins the two domains. A small C-terminal subdomain (CTSD; residues 379–407) extends the ErfK/YbiS/YhnG fold. In this subdomain, Trp394 and two Trp residues of the C-terminal helix a3 (398 and 401) make a zipper-like interaction with the IgD N-terminal domain that fixes | + | A short linker (residues 251–253) joins the two domains. A small <scene name='81/817533/Sub-domain/2'>C-terminal subdomain</scene> (CTSD; residues 379–407) extends the ErfK/YbiS/YhnG fold. In this subdomain, Trp394 and two Trp residues of the C-terminal helix a3 (398 and 401) make a zipper-like interaction with the IgD N-terminal domain that fixes the relative orientation of the two domains. |
| - | the relative orientation of the two domains. | + | |
== Active site == | == Active site == | ||
Revision as of 00:16, 17 June 2019
Overview
| |||||||||||