This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Insulin-Degrading Enzyme
From Proteopedia
(Difference between revisions)
| Line 7: | Line 7: | ||
IDE could help creating IDE-based therapies to control cerebral Aβ and blood sugar concentrations thanks to the understanding of its molecular mechanisms. | IDE could help creating IDE-based therapies to control cerebral Aβ and blood sugar concentrations thanks to the understanding of its molecular mechanisms. | ||
IDE is an unusual enzyme because of its high affinity for substrates that are widely diverse in sequence and structure. IDE prefers to degrade <6 kDa bioactive peptides such as insulin, Aβ, glucagon, atrial natriuretic peptides or transforming growth factor α. Paradoxally, even though IDE targets a broad range of substrates, it shows a remarkable capacity to selectively cleave some peptides without degrading related family members. | IDE is an unusual enzyme because of its high affinity for substrates that are widely diverse in sequence and structure. IDE prefers to degrade <6 kDa bioactive peptides such as insulin, Aβ, glucagon, atrial natriuretic peptides or transforming growth factor α. Paradoxally, even though IDE targets a broad range of substrates, it shows a remarkable capacity to selectively cleave some peptides without degrading related family members. | ||
| + | |||
| + | See [[Human Insulin Degrading Enzyme (Hebrew)]]. | ||
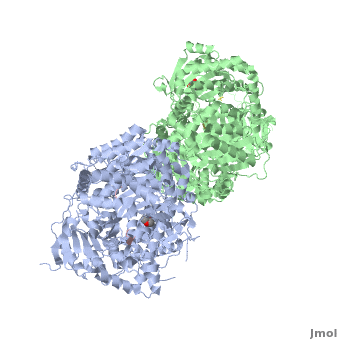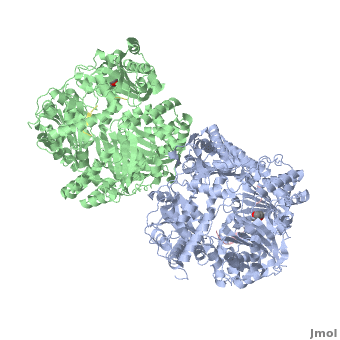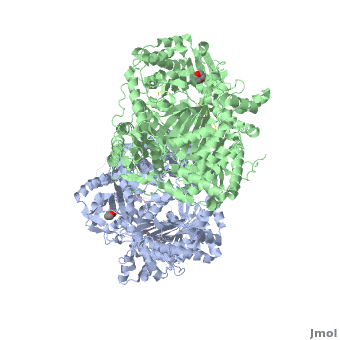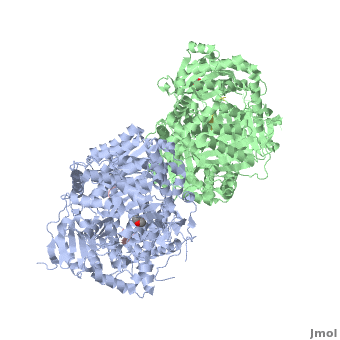
== How are the structure and the functions of IDE related? == | == How are the structure and the functions of IDE related? == | ||
Current revision
| |||||||||||
References
- Im H, Manolopoulou M, Malito E, Shen Y, Zhao J, Neant-Fery M, Sun CY, Meredith SC, Sisodia SS, Leissring MA, Tang WJ. Structure of substrate-free human insulin-degrading enzyme (IDE) and biophysical analysis of ATP-induced conformational switch of IDE. J Biol Chem. 2007 Aug 31;282(35):25453-63. Epub 2007 Jul 5. PMID:17613531 doi:10.1074/jbc.M701590200
- Shen Y, Joachimiak A, Rosner MR, Tang WJ. Structures of human insulin-degrading enzyme reveal a new substrate recognition mechanism. Nature. 2006 Oct 19;443(7113):870-4. Epub 2006 Oct 11. PMID:17051221 doi:10.1038/nature05143
- Li P, Kuo WL, Yousef M, Rosner MR, Tang WJ. The C-terminal domain of human insulin degrading enzyme is required for dimerization and substrate recognition. Biochem Biophys Res Commun. 2006 May 19;343(4):1032-7. Epub 2006 Mar 22. PMID:16574064 doi:10.1016/j.bbrc.2006.03.083
- ↑ Chen KC, Chang SS, Tsai FJ, Chen CY. Han ethnicity-specific type 2 diabetic treatment from traditional Chinese medicine? J Biomol Struct Dyn. 2012 Nov 12. PMID:23146021 doi:10.1080/07391102.2012.732340
- ↑ Leissring MA, Malito E, Hedouin S, Reinstatler L, Sahara T, Abdul-Hay SO, Choudhry S, Maharvi GM, Fauq AH, Huzarska M, May PS, Choi S, Logan TP, Turk BE, Cantley LC, Manolopoulou M, Tang WJ, Stein RL, Cuny GD, Selkoe DJ. Designed inhibitors of insulin-degrading enzyme regulate the catabolism and activity of insulin. PLoS One. 2010 May 7;5(5):e10504. PMID:20498699 doi:10.1371/journal.pone.0010504
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Élodie Weider, Karsten Theis, Joel L. Sussman, Jaime Prilusky