This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
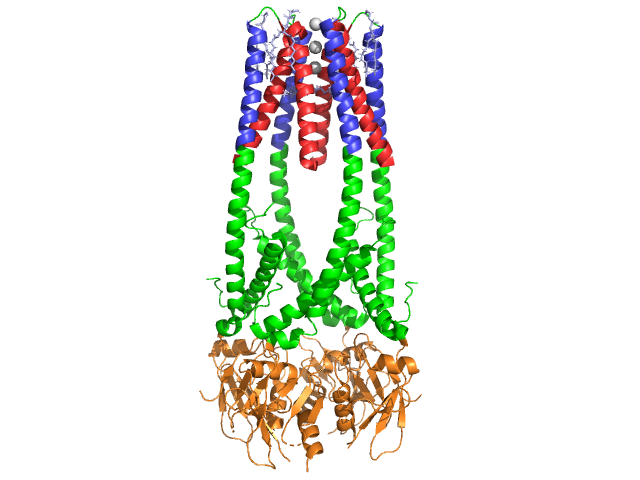
Lipase
From Proteopedia
(Difference between revisions)
| Line 66: | Line 66: | ||
</StructureSection> | </StructureSection> | ||
| - | == 3D Structures of Lipase == | ||
| - | |||
| - | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | ||
| - | {{#tree:id=OrganizedByTopic|openlevels=0| | ||
| - | |||
| - | *Eukaryote lipase: | ||
| - | |||
| - | **[[1hpl]] – hLip – horse <br /> | ||
| - | **[[1hlg]] – hLip – human - gastric<br /> | ||
| - | **[[1jmy]] – hBSSL <br /> | ||
| - | **[[1akn]] – cBSSL – cattle <br /> | ||
| - | **[[2bce]] - cBSSL (mutant) <br /> | ||
| - | **[[1f6w]] - cBSSL – catalytic domain<br /> | ||
| - | **[[3o0d]] – Lip – ''Yarrowia lipolytica''<br /> | ||
| - | **[[4jei]] – YlLip (mutant)<br /> | ||
| - | **[[1gpl]] – Lip – Guinea pig<br /> | ||
| - | **[[3zpx]] – Lip – ''Ustilago maydis''<br /> | ||
| - | **[[6a0w]] – Lip catalytic domain – bread mold<br /> | ||
| - | **[[4zrd]], [[4zre]] – Lip1 (mutant) – dandruff fungus<br /> | ||
| - | |||
| - | *Prokaryote lipase: | ||
| - | |||
| - | **[[1llf]] – Lip – ''Candida cylindracea''<br /> | ||
| - | **[[3g7n]] – Lip - ''Penicillium expansum''<br /> | ||
| - | **[[1tia]] - Lip – ''Penicillium camemberti''<br /> | ||
| - | **[[2hih]] – Lip – ''Staphylococcus hyicus''<br /> | ||
| - | **[[2fx5]] – Lip – ''Pseudomonas mendocina''<br /> | ||
| - | **[[1yzf]] – Lip – ''Enterococcus faecalis''<br /> | ||
| - | **[[1dt3]], [[1dt5]], [[1dte]], [[1du4]], [[1ein]], [[1tib]], [[4dyh]], [[4ea6]], [[4flf]], [[4gbg]], [[4gwl]], [[4zgb]], [[6hw1]] - TlLip - ''Thermomyces lanuginose''<br /> | ||
| - | **[[5ap9]] – TlLip (mutant)<br /> | ||
| - | **[[1jfr]] – Lip – ''Streptomyces exfoliates''<br /> | ||
| - | **[[5mal]] – Lip – ''Streptomyces rimosus''<br /> | ||
| - | **[[2lip]] – BcLip – open state<br /> | ||
| - | **[[1cvl]] – Lip – ''Chromobacterium viscosum''<br /> | ||
| - | **[[1tic]] - Lip – ''Rhizopus oryzae''<br /> | ||
| - | **[[3tgl]], [[4tgl]], [[1tgl]], [[6qpp]] – RmLip – ''Rhyzomucor miehei''<br /> | ||
| - | **[[6qpr]] – RmLip (mutant)<br /> | ||
| - | **[[2zvd]] – PsLip - ''Pseudomonas sp.'' – open state<br /> | ||
| - | **[[2z8x]] - PsLip – extracellular<br /> | ||
| - | **[[5xpx]] – PsLip residues 1-388<br /> | ||
| - | **[[2zj6]], [[2zj7]] – PsLip (mutant) <br /> | ||
| - | **[[2z8z]] – PsLip(mutant) – closed state<br /> | ||
| - | **[[3lip]], [[3a6z]] - Lip - ''Pseudomonas cepacia'' – open state<br /> | ||
| - | **[[1qge]], [[1tah]] – Lip – ''Pseudomonas glumae''<br /> | ||
| - | **[[2w22]], [[6a12]] – GtLip – ''Geobacillus thermocatenulatus''<br /> | ||
| - | **[[5ce5]] – GtLip (mutant)<br /> | ||
| - | **[[1ji3]], [[1ku0]], [[4fmp]], [[4x6u]] – BstLip – ''Bacillus stearothermophilus''<br /> | ||
| - | **[[4x71]], [[4x7b]], [[4x85]], [[6s3g]], [[6s3j]], [[6s3v]], [[6fz1]], [[6fz7]], [[6fz8]], [[6fz9]], [[6fza]], [[6fzc]], [[6fzd]] – BstLip (mutant)<br /> | ||
| - | **[[1ah7]] - Lip – ''Bacillus cereus''<br /> | ||
| - | **[[2ory]] – Lip – ''Photobacterium lypoliticum''<br /> | ||
| - | **[[2z5g]], [[2dsn]] – GzLip T1 – ''Geobacillus zalihae''<br /> | ||
| - | **[[3umj]] – GzLip (mutant)<br /> | ||
| - | **[[3p94]] – Lip – ''Parabacteroides distasonis''<br /> | ||
| - | **[[3ngm]] – Lip – ''Gibberella zeae''<br /> | ||
| - | **[[3auk]] - Lip – ''Geobacillus''<br /> | ||
| - | **[[3uue]], [[3uuf]] – Lip – ''Malassezia globosa''<br /> | ||
| - | **[[4hs9]] – Lip – ''Proteus mirabilis''<br /> | ||
| - | **[[4opm]] – Lip – ''Acinetobacter baumannii''<br /> | ||
| - | **[[5ah0]] – Lip – ''Pelosinus fermentans''<br /> | ||
| - | **[[5ah1]] – Lip – ''Clostridium botulinum''<br /> | ||
| - | **[[5ch8]] – Lip (mutant) – ''Penicillium cyclopium''<br /> | ||
| - | |||
| - | *Bacterial lipase A | ||
| - | |||
| - | **[[3guu]] – CaLipA – ''Candida Antarctica''<br /> | ||
| - | **[[2veo]] – CaLipA – closed state<br /> | ||
| - | **[[2qua]], [[2qub]] – SmLipA – ''Serratia marcescens''<br /> | ||
| - | **[[2qxt]], [[2qxu]], [[1isp]], [[1i6w]], [[5ct4]], [[5ct5]], [[5ct6]], [[5cri]] - BsLipA – ''Bacillus subtilis''<br /> | ||
| - | **[[3d2a]], [[3d2b]], [[3d2c]], [[1t2n]], [[1t4m]], [[5ct8]], [[5ct9]], [[5cta]], [[5cur]], [[3qzu]], [[3qmm]] - BsLipA (mutant) <br /> | ||
| - | **[[1r4z]] – BsLipA+Rc-IPG-phosphonate<br /> | ||
| - | **[[1r50]] – BsLipA +Sc-IPG-phosphonate<br /> | ||
| - | |||
| - | *Bacterial lipase B | ||
| - | |||
| - | **[[4zv7]], [[3w9b]], [[5a6v]], [[5a71]], [[4k6g]], [[1tca]], [[1tcb]], [[1tcc]], [[1lbs]], [[1lbt]] – CaLipB <br /> | ||
| - | **[[3icv]], [[4k5q]], [[5k6h]], [[5k6k]] – CaLipB (mutant) <br /> | ||
| - | **[[5gv5]] - CaLipB + phosphonate<br /> | ||
| - | **[[3icw]] – CaLipB (mutant) + phosphonate<br /> | ||
| - | **[[5x7k]] – SmLipB NBD <br /> | ||
| - | **[[6isp]], [[6isq]], [[6isr]] – LipB (mutant) – ''Pseudozyma antarctica''<br /> | ||
| - | **[[6idy]] - AfLipB - ''Aspergillus fumigatus''<br /> | ||
| - | |||
| - | *Bacterial lipase C | ||
| - | |||
| - | **[[5nen]] – SmLipC residues 1-443 <br /> | ||
| - | |||
| - | *Lipase/colipase complexes. The colipase is a co-enzyme whose binding to lipase optimizes the enzymatic activity | ||
| - | |||
| - | **[[1n8s]] – hLip+colipase II<br /> | ||
| - | **[[1eth]], [[1lpa]] - pLip+colipase II - pig<br /> | ||
| - | |||
| - | *Hormone-sensitive-lipases (LIPE) hydrolyze the first fatty acid of the triacylglycerol substrate | ||
| - | |||
| - | **[[3k6k]] – EstE7(LIPE) – metagenome library<br /> | ||
| - | **[[3fak]], [[3dnm]] – EstE5(LIPE) – metagenome library<br /> | ||
| - | **[[1evq]] – AaEst2(LIPE) – ''Alicyclobacillus acidocaldarius''<br /> | ||
| - | **[[1u4n]] – AaEst2(LIPE) (mutant) <br /> | ||
| - | |||
| - | *Putative lipases; Proteins with unknown function but structural similarity to lipase obtained in structural genomics projects. | ||
| - | |||
| - | **[[2rau]] - Lip – ''Sulfolobus solfataricus''<br /> | ||
| - | **[[3bxp]], [[3d3n]] - Lip – ''Lactobacillus plantarum''<br /> | ||
| - | **[[3e0x]] - Lip – ''Clostridium acetobutylicum''<br /> | ||
| - | **[[1z8h]] – Lip – ''Nostoc sp.'' PCC 712<br /> | ||
| - | **[[1vj3]] - Lip – ''Nostoc sp.'' <br /> | ||
| - | **[[3bzw]] – Lip - ''Bacteroides thetaiotaomicron''<br /> | ||
| - | **[[2pbl]] – Lip - ''Silicibacter'' | ||
| - | |||
| - | *Lipase + inhibitors | ||
| - | |||
| - | **[[3l1h]] – EstE5(LIPE)+FeCl3 – noninvasive inhibitor<br /> | ||
| - | **[[3l1i]], [[3l1j]] - EstE5(LIPE)+CuSO4 – noninvasive inhibitor<br /> | ||
| - | **[[3lij]] - EstE5(LIPE)+ZnSO4– noninvasive inhibitor<br /> | ||
| - | **[[3h18]], [[3h17]] - EstE5 (LIPE)+PMSF <br /> | ||
| - | **[[3h19]], [[3h1b]], [[3h1a]] – EstE5 (LIPE)+methyl alcohol<br /> | ||
| - | **[[3h1a]] – EstE5 SLIPE)+ethyl alcohol<br /> | ||
| - | **[[3h19]] – EstE5 SLIPE)+isopropyl alcohol<br /> | ||
| - | **[[3g9t]], [[3g9u]] - EstE5 (HSLIPE)+p-nitrophenyl butyrate<br /> | ||
| - | **[[3g9z]] - EstE5 (LIPE) +p-nitrophenyl caprylate<br /> | ||
| - | **[[2nw6]] – BcLip+ S inhibitor <br /> | ||
| - | **[[4lip]], [[5lip]], [[1r4z]], [[1r50]] – BcLip+ Rc-(Rp,Sp)-1,2-dioctylcarbamoyl-glycero-3-O-phosphonate<br /> | ||
| - | **[[1k8q]] - Lip+phosphonate – dog <br /> | ||
| - | **[[1ex9]] – Lip+Rc-(Rp,Sp)-1,2-dioctylcarbamoyl-glycero-3-O-phosphonate – ''Pseudomonas aeruginosa'' <br /> | ||
| - | **[[5tgl]] – RmLip+N-hexyl-phosphonate <br /> | ||
| - | **[[1lpb]] – pLip + colipase+C11 alkyl phosphonate <br /> | ||
| - | **[[3a70]] – PsLip+diethyl phosphate<br /> | ||
| - | **[[4glb]] – TlLip + nitrobenzaldehyde<br /> | ||
| - | **[[4kjx]] - TlLip + nitrobenzaldehyde + lauric acid<br /> | ||
| - | **[[4n8s]] - TlLip + nitrobenzaldehyde + ethylacetoacetate<br /> | ||
| - | **[[4s0x]] – TlLip + lauric acid<br /> | ||
| - | |||
| - | *Lipase conjugated with analogs to its reaction intermediates | ||
| - | |||
| - | **[[1qz3]] – EaEst2(mutant) (LIPE)+hexadecanesulfonate <br /> | ||
| - | |||
| - | *Bile-salt activated lipase | ||
| - | |||
| - | **[[6h0t]] – hBAL (mutant) <br /> | ||
| - | **[[6h0v]], [[6h18]], [[6h19]], [[6h1a]] – hBAL (mutant) + nerve agent<br /> | ||
| - | **[[1aql]] – bBAL+taurocholate - bovine<br /> | ||
| - | |||
| - | *Monoacylglycerol lipase | ||
| - | |||
| - | **[[3hju]], [[3jw8]] - hMAGL <BR /> | ||
| - | **[[3jwe]], [[3pe6]], [[4uuq]], [[6ax1]], [[6bq0]] – hMAGL + inhibitor<br /> | ||
| - | **[[4uuq]] - hMAGLip + SAR <br /> | ||
| - | **[[5zun]] – hMAGL (mutant) + inhibitor<br /> | ||
| - | **[[3rm3]], [[4lhe]], [[5xks]] - BaMAGL – ''Bacillus'' <BR /> | ||
| - | **[[3rli]] – BaMAGL + PMSF<br /> | ||
| - | **[[4ke7]], [[4ke8]], [[4ke9]] – BaMAGL + ligand<br /> | ||
| - | **[[4ke6]], [[4kea]] – BaMAGL (mutant) + ligand<br /> | ||
| - | **[[4zwn]] – yMAGL – yeast<br /> | ||
| - | **[[4zxf]] – yMAGLip + substrate analog<br /> | ||
| - | **[[6eic]] – MAGLip – ''Mycobacterium tuberculosis''<br /> | ||
| - | **[[5xk2]] – MAGLip – ''Aespergillus oryzae''<br /> | ||
| - | |||
| - | *Lipase with substrate bound at active site | ||
| - | |||
| - | **[[2zyh]] – AfLip (mutant)+fatty acid – ''Archaeoglobus fulgidus''<br /> | ||
| - | **[[2zyi]], [[2zyr]], [[2zys]] - AfLip+fatty acid+ ion <br /> | ||
| - | **[[1gt6]], [[4ghw]], [[4gi1]] – TlLip+ fatty acid - lipid ligand<br /> | ||
| - | |||
| - | *Lipase conjugated to transition-state analogs showing the binding mode of the enzyme catalysis | ||
| - | |||
| - | **[[1ys1]] – BhLip+hexylphosphonic acid (R) 2-methyl-3-phenylpropyl ester <br /> | ||
| - | **[[1ys2]] – BhLip+hexylphosphonic acid (S) 2-methyl-3-phenylpropyl ester<br /> | ||
| - | **[[1hqd]] – Lip+1-phenoxy-2-acrtoxy butane – ''Pseudomonas cepacia'' <br /> | ||
| - | |||
| - | *Lipase+lipase chaperone | ||
| - | |||
| - | **[[2es4]] – Lip+lipase chaperone C-terminal - ''Burkholderia glumae'' | ||
| - | |||
| - | *Lipase 2 or esterase/lipase | ||
| - | |||
| - | **[[3v9a]],[[3g9t]], [[3g9u]], [[3g9z]], [[3h19]], [[3h1a]], [[3h1b]], [[3l1h]], [[3l1i]], [[3l1j]], [[3fak]], [[3h17]], [[3h18]], [[3dnm]], [[3k6k]] - E/L - uncultured bacteria<br /> | ||
| - | **[[3w9b]] - CaE/L <br /> | ||
| - | **[[4v2i]] - E/L - ''Thalassospira''<br /> | ||
| - | **[[4n5h]] - LrE/L (mutant) - ''Lactobacillus rhamnosus''<br /> | ||
| - | **[[4n5i]] - LrE/L + inhibitor <br /> | ||
| - | **[[4ouk]] - LrE/L (mutant) + inhibitor <br /> | ||
| - | **[[4bzz]], [[4bzw]] - LpE/L <br /> | ||
| - | **[[6gup]] - AfE/L <br /> | ||
| - | **[[1lgy]] – Lip2 – ''Rhizopus niveus''<br /> | ||
| - | **[[1gz7]] - CrLip2 <br /> | ||
| - | **[[4jei]] – Lip2 – ''Yarrowia lipolytica''<br /> | ||
| - | **[[1thg]] – Lip2 – ''Geotrichum candidum''<br /> | ||
| - | }} | ||
==References== | ==References== | ||
<references /> | <references /> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Revision as of 09:58, 22 October 2019
| |||||||||||
References
- ↑ [1] 1HPL PDB SUM
- ↑ [2] A cross-linked complex between horse pancreatic lipase and colipase
- ↑ [3] History of Lipids
- ↑ [4] 1HPL PDB
- ↑ http://www.pdb.org/pdb/explore/explore.do?structureId=1HPL
- ↑ http://www.pdb.org/pdb/explore/remediatedSequence.do?structureId=1HPL
- ↑ http://www.springerlink.com/content/g5h1613440115701/fulltext.pdf
- ↑ Fundamentals of Biochemistry...
- ↑ Thomas, A. etc. "Role of the Lid Hydrophobicity Pattern in Pancreatic Lipase Activity", The Journal of Biological Chemistry, 2005 September 22; 270 (48): 40074-40083.
- ↑ "Colipase". Wikipedia: The Free Encyclopedia. 5 July 2011 [5]
- ↑ "Colipase Residues..."
- ↑ Fundamentals of Biochemistry...
- ↑ Crandall,W., Lowe, M. "Colipase Residues Glu64 and Arg65 Are Essential for Normal Lipase-mediated Fat Digestion in the Presence of Bile Salt Micelles" Journal of Biological Chemistry, 2001, (276) 12505-12512
- ↑ van Tilbeurgh H, etc."Structure of the pancreatic lipase-procolipase complex", 1992 Sep 10;359(6391):159-62. PMID:1522902.[6]
- ↑ http://www.pdb.org/pdb/explore/explore.do?structureId=1ETH
- ↑ http://www.nature.com/nature/journal/v362/n6423/abs/362814a0.html
- ↑ Sussman JL, Harel M, Frolow F, Oefner C, Goldman A, Toker L, Silman I. Atomic structure of acetylcholinesterase from Torpedo californica: a prototypic acetylcholine-binding protein. Science. 1991 Aug 23;253(5022):872-9. PMID:1678899
- ↑ Ollis DL, Cheah E, Cygler M, Dijkstra B, Frolow F, Franken SM, Harel M, Remington SJ, Silman I, Schrag J, et al.. The alpha/beta hydrolase fold. Protein Eng. 1992 Apr;5(3):197-211. PMID:1409539
- ↑ Bourne Y, Martinez C, Kerfelec B, Lombardo D, Chapus C, Cambillau C. Horse pancreatic lipase. The crystal structure refined at 2.3 A resolution. J Mol Biol. 1994 May 20;238(5):709-32. PMID:8182745 doi:http://dx.doi.org/10.1006/jmbi.1994.1331
- ↑ [7] 1LPB PDB SUM
- ↑ "Pancreatic lipase". Wikipedia: The Free Encyclopedia. 7 Nov 2011 [8]
- ↑ Kordik, C., Reitz, A. "Pharmacological Treatment of Obesity: Therapeutic Strategies" Journal of Medicinal Chemistry, 1999 (42).
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Quinn R. Murray, Natalie Ziegler, Stephanie Schell, David Canner, Alexander Berchansky, Katelyn Clark, Eric Martz, Leben Tadesse, Joel L. Sussman, Eran Hodis