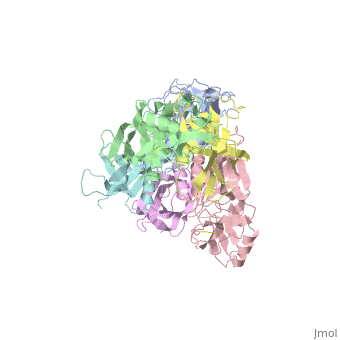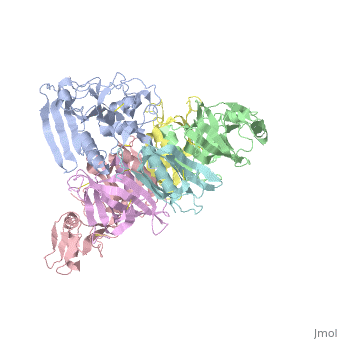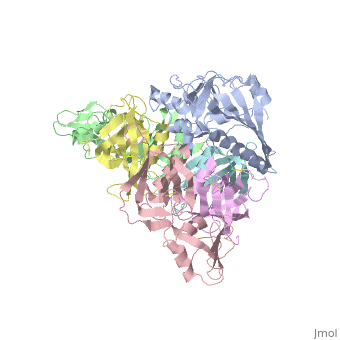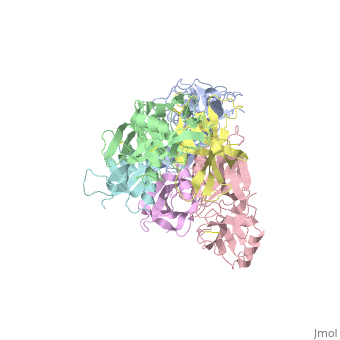This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1prt
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
<StructureSection load='1prt' size='340' side='right'caption='[[1prt]], [[Resolution|resolution]] 2.90Å' scene=''> | <StructureSection load='1prt' size='340' side='right'caption='[[1prt]], [[Resolution|resolution]] 2.90Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1prt]] is a 12 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[1prt]] is a 12 chain structure with sequence from [https://en.wikipedia.org/wiki/"bacterium_tussis-convulsivae"_lehmann_and_neumann_1927 "bacterium tussis-convulsivae" lehmann and neumann 1927]. The September 2005 RCSB PDB [https://pdb.rcsb.org/pdb/static.do?p=education_discussion/molecule_of_the_month/index.html Molecule of the Month] feature on ''Cholera Toxin'' by David S. Goodsell is [https://dx.doi.org/10.2210/rcsb_pdb/mom_2005_9 10.2210/rcsb_pdb/mom_2005_9]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1PRT OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1PRT FirstGlance]. <br> |
| - | </td></tr><tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | </td></tr><tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1prt FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1prt OCA], [https://pdbe.org/1prt PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1prt RCSB], [https://www.ebi.ac.uk/pdbsum/1prt PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1prt ProSAT]</span></td></tr> |
</table> | </table> | ||
== Function == | == Function == | ||
| - | [[ | + | [[https://www.uniprot.org/uniprot/TOX4_BORPE TOX4_BORPE]] PTX oligomer B binds to receptors on the eukaryotic cell surface and facilitates the translocation of the toxic subunit across the cell membrane. [[https://www.uniprot.org/uniprot/TOX1_BORPE TOX1_BORPE]] S1 is an NAD-dependent ADP-ribosyltransferase, which plays a crucial role in the pathogenesis of B.pertussis causing disruption of normal host cellular regulation. It catalyzes the ADP-ribosylation of a cysteine in the alpha subunit of host heterotrimeric G proteins. In the absence of G proteins it also catalyzes the cleavage of NAD(+) into ADP-ribose and nicotinamide. It irreversibly uncouples the G-alpha GTP-binding proteins from their membrane receptors. [[https://www.uniprot.org/uniprot/TOX3_BORPE TOX3_BORPE]] PTX oligomer B binds to receptors on the eukaryotic cell surface and facilitates the translocation of the toxic subunit across the cell membrane. [[https://www.uniprot.org/uniprot/TOX2_BORPE TOX2_BORPE]] PTX oligomer B binds to receptors on the eukaryotic cell surface and facilitates the translocation of the toxic subunit across the cell membrane. [[https://www.uniprot.org/uniprot/TOX5_BORPE TOX5_BORPE]] PTX oligomer B binds to receptors on the eukaryotic cell surface and facilitates the translocation of the toxic subunit across the cell membrane. |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
Revision as of 09:33, 8 September 2021
THE CRYSTAL STRUCTURE OF PERTUSSIS TOXIN
| |||||||||||