This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1558
From Proteopedia
(Difference between revisions)
| Line 19: | Line 19: | ||
The <scene name='82/823082/Catalytic_triad/2'>Catalytic Triad</scene> of this protein is primarily made of the amino acids that are the main factor in catalysis. The 3 amino acids are Phe-59, Tyr101, and Lys-134. The <scene name='82/823082/Colored_secondary/1'>secondary and terteriary structure</scene> is a fold of LsdA of a seven-bladed -propeller, typical of the carotenoid cleavage oxygenates (CCO's), which usually catalyze the oxidative cleavage of a double bond in carotenoids. | The <scene name='82/823082/Catalytic_triad/2'>Catalytic Triad</scene> of this protein is primarily made of the amino acids that are the main factor in catalysis. The 3 amino acids are Phe-59, Tyr101, and Lys-134. The <scene name='82/823082/Colored_secondary/1'>secondary and terteriary structure</scene> is a fold of LsdA of a seven-bladed -propeller, typical of the carotenoid cleavage oxygenates (CCO's), which usually catalyze the oxidative cleavage of a double bond in carotenoids. | ||
| - | The structures also consist of α-helices and ß-sheets. The <scene name='82/823082/Hydrogen_bonding/1'>active site</scene> occurs at the center of the propeller and contains an Fe2+. The <scene name='82/823082/Hydrophobicity_whole_protein/1'> | + | The structures also consist of α-helices and ß-sheets. The <scene name='82/823082/Hydrogen_bonding/1'>active site</scene> occurs at the center of the propeller and contains an Fe2+. The <scene name='82/823082/Hydrophobicity_whole_protein/1'>hydrophobicity</scene> and <scene name='82/823082/Spacefill_whole_protein/1'>spacefill</scene> view of the ligand in the protein, which shows that both hydrophilic and hydrophobic residues are important to the ligand in the binding site. |
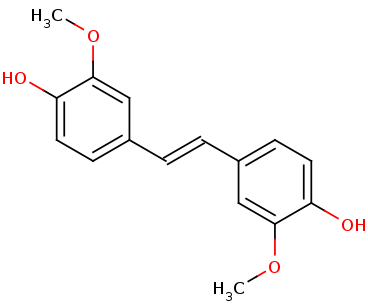
The space fill view allows us to see the different binding sites for this protein, the binding site is allosteric. The binding site for this protein has a very restrictive accessibility. The <scene name='82/823082/Ligand_closeup/2'>ligand</scene> for this protein is called NSL. | The space fill view allows us to see the different binding sites for this protein, the binding site is allosteric. The binding site for this protein has a very restrictive accessibility. The <scene name='82/823082/Ligand_closeup/2'>ligand</scene> for this protein is called NSL. | ||
Revision as of 02:39, 9 December 2019
| This Sandbox is Reserved from Aug 26 through Dec 12, 2019 for use in the course CHEM 351 Biochemistry taught by Bonnie_Hall at the Grand View University, Des Moines, USA. This reservation includes Sandbox Reserved 1556 through Sandbox Reserved 1575. |
To get started:
More help: Help:Editing |
Lignostilbene-α,β-dioxygenase A (LsdA) Catalyzation
| |||||||||||
References
Kuatsjah, Eugene, et al. “Identification of Functionally Important Residues and Structural Features in a Bacterial Lignostilbene Dioxygenase.” Journal of Biological Chemistry, vol. 294, no. 35, 2019, pp. 12911–12920., doi:10.1074/jbc.ra119.009428.
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644