This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1607
From Proteopedia
(Difference between revisions)
(New page: {{Sandbox_Reserved_CH462_Biochemistry_II}}<!-- PLEASE ADD YOUR CONTENT BELOW HERE --> ==Your Heading Here (maybe something like 'Structure')== <StructureSection load='1stp' size='340' side...) |
|||
| Line 1: | Line 1: | ||
| - | + | =Mitochondrial Calcium Uniporter= | |
| - | + | <StructureSection load='6DT0' size='350' frame='true' side='right' caption='Calcium Uniporter 6DT0' scene=’’> | |
| - | <StructureSection load=' | + | This is a default text for your page '''Elizabeth M. Ratz/Sandbox 1'''. Click above on '''edit this page''' to modify. Be careful with the < and > signs. |
| - | This is a default text for your page ''''''. Click above on '''edit this page''' to modify. Be careful with the < and > signs. | + | |
You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> or to the article describing Jmol <ref>PMID:21638687</ref> to the rescue. | You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> or to the article describing Jmol <ref>PMID:21638687</ref> to the rescue. | ||
| + | |||
| + | ==Introduction== | ||
| + | |||
| + | <scene name='83/837222/Selectivity_filter/3'>Selectivity Filter</scene> | ||
| + | |||
| + | ==Structure== | ||
| + | |||
| + | ===Selectivity Filter=== | ||
| + | |||
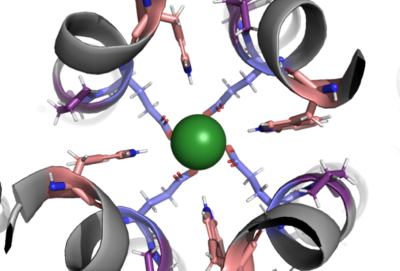
| + | The selectivity filter contains Glu358, Trp354, and Pro359 to allow calcium to pass through the uniporter. The carboxylate oxygen of the Glu side chains draw in the positive calcium ion. The diameter of the carboxyl ring is 5Å, allowing only a dehydrated Ca ion to bind. Trp38, which is directly next to the Glu residues, stabilizes the carbonyl side chains through hydrogen bonding and anion pi interactions. These Trp residues also form stacking interactions with Pro359, which orientate the Glu carboxyl side chains towards the middle of the pore to interact with Ca ions. | ||
| + | |||
| + | [[Image:Selectivity_filter.png|400 px|right|thumb|Figure 1]] | ||
| + | |||
| + | [http://www.rcsb.org/structure/6DT0 Calcium Uniporter Structure] | ||
| + | |||
| + | <ref name=”Yoo”>PMID:29954988</ref> | ||
== Function == | == Function == | ||
| - | == Disease == | + | ===Inhibitors=== |
| + | [[Image:Screen Shot 2020-03-30 at 4.37.32 PM.png|400 px|left|thumb|Figure 2]] | ||
| + | |||
| + | == Disease Links== | ||
| + | |||
| + | ===Types 1 and 2 Diabetes=== | ||
| + | |||
| + | Pancreatic beta cells circulate insulin through the body. Glucose initiates signals allowing these cells to break down sugar and release insulin, which is all stimulated by mitochondrial energy metabolism. Calcium homeostasis plays a fundamental role in ATP production supplying energy to this process. Defects in homeostasis of calcium like chronic calcium depletion, caused by leaky Ryanodine receptors, causes types 1 and 2 diabetes through failure of this mechanism. Treatment involves targeting the MCU and MICU1 to open the calcium channel and allow more uptake of calcium ions into the mitochondria. | ||
| + | |||
| + | ===Heart Failure=== | ||
| + | |||
| + | Calcium overload in the mitochondria of cardiac cells lead to apoptotic cardiac cell death. This large amount of cell death | ||
== Relevance == | == Relevance == | ||
== Structural highlights == | == Structural highlights == | ||
| + | |||
| + | == Student Contributors == | ||
This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | ||
Revision as of 20:50, 30 March 2020
Mitochondrial Calcium Uniporter
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Yoo J, Wu M, Yin Y, Herzik MA Jr, Lander GC, Lee SY. Cryo-EM structure of a mitochondrial calcium uniporter. Science. 2018 Jun 28. pii: science.aar4056. doi: 10.1126/science.aar4056. PMID:29954988 doi:http://dx.doi.org/10.1126/science.aar4056