This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Ivan E. Wang/Sandbox Reserved 895
From Proteopedia
| Line 1: | Line 1: | ||
= Introduction to the RPE65 = | = Introduction to the RPE65 = | ||
<StructureSection load='4rsc' size='500' side='right' caption='Figure 1: Emixustat and palmitate bound in the active site of RPE65' scene=''> | <StructureSection load='4rsc' size='500' side='right' caption='Figure 1: Emixustat and palmitate bound in the active site of RPE65' scene=''> | ||
| + | |||
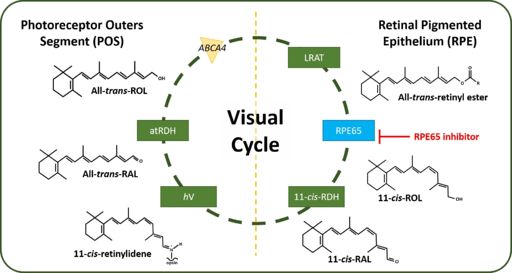
Retinal pigment epithelium 65 (RPE65) also known as retinoid isomerohydrolase (IMH) is a 65 kDa enzyme located within the human retinal pigment epithelium (hRPE) cells. RPE65 is exclusively expressed within hRPE cells and is not expressed anywhere else within the body which makes RPE65 a useful drug target. PRE65 is responsible for the chemical conversion of ''all-trans''-retinyl ester to ''11-cis''-retinol, the rate limiting step, in the canonical visual cycle. | Retinal pigment epithelium 65 (RPE65) also known as retinoid isomerohydrolase (IMH) is a 65 kDa enzyme located within the human retinal pigment epithelium (hRPE) cells. RPE65 is exclusively expressed within hRPE cells and is not expressed anywhere else within the body which makes RPE65 a useful drug target. PRE65 is responsible for the chemical conversion of ''all-trans''-retinyl ester to ''11-cis''-retinol, the rate limiting step, in the canonical visual cycle. | ||
| Line 6: | Line 7: | ||
= Structure and Activity = | = Structure and Activity = | ||
| - | == Structural Analysis of RPE65 == | + | == Classification and Structural Analysis of RPE65 == |
| - | ( | + | |
| + | Classification nomenclature for RPE65 is as follow according to <insert classification type> | ||
| + | |||
| + | Retinal pigment epithelium 65 (RPE65) resembles a 7-bladed propeller with two deep binding pockets that contains a lipophilic binding pocket (which binds to endogenous palmitoyl acid) as well as an iron(II) atom stabilized by 5 histidine residues. (<insert histidine residue positions>) | ||
| + | |||
== Enzymatic Activity of RPE65 == | == Enzymatic Activity of RPE65 == | ||
Revision as of 18:29, 25 February 2020
Contents |
Introduction to the RPE65
| |||||||||||
Structure and Activity
Classification and Structural Analysis of RPE65
Classification nomenclature for RPE65 is as follow according to <insert classification type>
Retinal pigment epithelium 65 (RPE65) resembles a 7-bladed propeller with two deep binding pockets that contains a lipophilic binding pocket (which binds to endogenous palmitoyl acid) as well as an iron(II) atom stabilized by 5 histidine residues. (<insert histidine residue positions>)
Enzymatic Activity of RPE65
(PLACEHOLDER)
Protein-Ligand Interaction
Endogenous Ligand
(PLACEHOLDER)
Exogenous Ligand
(R)-Emixustat (ACU-4429)
| |||||||||||
(S)-Emixustat
| |||||||||||
R/S Enantiomers Differences
(PLACEHOLDER)
Disease Implications
(PLACEHOLDER)
Medical Relevance
(PLACEHOLDER)
This is a sample scene created with SAT to by Group, and another to make of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes.
References
- ↑ Shin Y, Moiseyev G, Petrukhin K, Cioffi CL, Muthuraman P, Takahashi Y, Ma JX. A novel RPE65 inhibitor CU239 suppresses visual cycle and prevents retinal degeneration. Biochim Biophys Acta Mol Basis Dis. 2018 Jul;1864(7):2420-2429. doi:, 10.1016/j.bbadis.2018.04.014. Epub 2018 Apr 21. PMID:29684583 doi:http://dx.doi.org/10.1016/j.bbadis.2018.04.014
