Introduction
Bd oxidase is an integral membrane protein that catalyzes the reduction of O2 to 2H20 using quinol as the reducing substrate. The reaction is electrogenic but is not coupled to a proton pump it instead uses internal water molecules to provide the protons needed for the reduction reaction. It plays a key role in protecting the organism from high oxidative stress (ie. preventing free radicals in intracellular space in prokaryotes, more specifically gram-negative heterotrophs).
There are two main types of respiratory cytochrome oxidases: the heme/copper oxidases, and the heme-only cytochrome bd quinol oxidase, which is what bd oxidase falls under. Heme-only cytochrome bd quinol oxidases are associated with microaerobic dioxygen respiration, and they have a high affinity for oxygen.
Structure
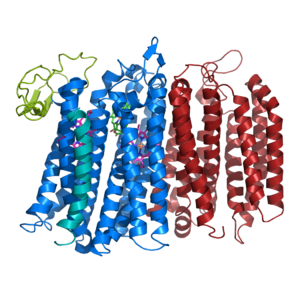
Figure 1. bd oxidase; two structurally similar transmembrane helices in blue and red; CydX subunit in teal; Q loop in lime green.
The overall structure contains 19 transmembrane helices that are arranged in a nearly oval shape (Fig 1.). The protein contains two structurally similar subunits each containing nine helices (blue and red) and one smaller subunit, CydX, with one transmembrane helix. The subunits are interacting using hydrophobic residues and symmetry at the interfaces. The CydX subunit, whose function is not currently known, is positioned in the same way as CydS, which is found in E. coli bd oxidase. Due to its similar structure and position, it has been hypothesized to potentially stabilize heme b558 during potential structural rearrangements of the Q loop upon binding and oxidation of quinol. The Q loop is shown in lime green, and is a hydrophilic region above Cyd A. The lack of hydrogen bonding in this hydrophobic protein allows the protein to be flexible and go through a large conformational change for reduction of dioxygen.
Active Site
The active site for Bd Oxidase in Geobacillus thermodenitrificans is located in subunit Cyd A. The site consists of three iron hemes: Heme B558, Heme B595, and Heme D that are held together in a rigid triangular due to van der wals interactions. The length between each heme's central iron is relatively constant which serves to shuttle protons and electrons from one heme to another efficiently. It is suggested that Heme B558 acts as an electron acceptor to the extracellular side and Heme B559 acts as a proton acceptor on the intracellular side. It is then proposed that both heme B558 and B595 shuttle their respective ions directly to Heme D based on this being the shortest pathway (reference). Heme D is then suggested to be the oxygen binding site due to proximity and orientation to the exterior surface of the protein.
Potential Oxygen Entry Site
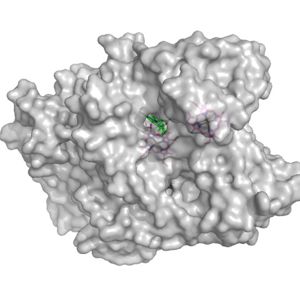
Figure 2. Surface of the potential oxygen entry site; Heme D shown in green
Heme D is the hypothesized spot for the oxygen to enter the protein. Heme D is directly connected to the protein surface on CydA and contains a solvent accessible substrate channel (Fig 2.)
Electron Source
An electron source is needed in order for the redox reaction of O₂ to occur. Bd oxidase uses the quinol molecule ubiquinone as an electron donor. The chemical structure of ubiquinone is shown in Fig. 3.

Figure 3. Chemical structure of ubiquinone.
Potential Proton Pathways
Disease
Relevance
Structural highlights
This is a sample scene created with SAT to by Group, and another to make of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes.



