This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Sumit Kamat/Sandbox Reserved 901
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
== Overview == | == Overview == | ||
| - | Histone-lysine N-methyltransferase 2A (KMT2A) known as | + | Histone-lysine N-methyltransferase 2A (KMT2A) known as mixed-lineage leukemia 1 (MLL-1) is an enzyme that in humans is encoded by the KMT2A gene. KMT2A gene is a histone methyltransferase which are histone modifying enzymes which catalyze the transfer of methyl groups to lysine and arginine residues of histone proteins. The KMT2A gene is a positive global regulator of gene transcription and comprises of transactivation domain 9aaTAD which is involved in the epigenetic maintenance of transcriptional memory <ref>PMID 1720549</ref>. The KMT2 family can mono-, di- and trimethylates histone H3K4. This family of enzymes is found within a macromolecular complex known as the COMPASS family and are highly conserved from yeast to human <ref>PMC3711870</ref>. |
| Line 16: | Line 16: | ||
]] | ]] | ||
| + | == Structural Analysis == | ||
| + | |||
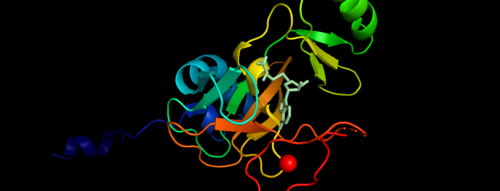
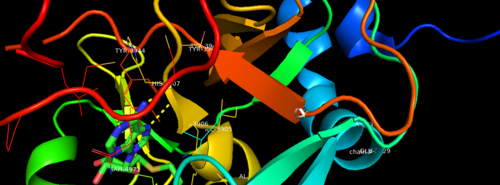
| + | MLL-1 is a 431-kDa protein to be a structural and functional homolog of the Drosophila trithorax (TRX) protein. Two domains are highly conserved between MLL and TRX consist of a carboxy-terminal SET (Su(var)3-9, enhancer-of-zeste, and trithorax) domain and internal plant homeodomain (PHD) fingers. Both domains are found in many chromatin-associated transcriptional regulators and are thought to function either directly in chromatin modification or as protein-protein interaction surfaces for the recruitment of chromatin-modifying machinery <ref> https://doi.org/10.1182/blood-2002-04-1015 </ref>. | ||
| Line 22: | Line 25: | ||
== Relevance == | == Relevance == | ||
| - | + | ||
This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | ||
Revision as of 17:50, 28 April 2020
Histone-lysine N-methyltransferase 2A KMT2A
| |||||||||||