This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
3frp
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
<StructureSection load='3frp' size='340' side='right'caption='[[3frp]], [[Resolution|resolution]] 2.61Å' scene=''> | <StructureSection load='3frp' size='340' side='right'caption='[[3frp]], [[Resolution|resolution]] 2.61Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[3frp]] is a 3 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[3frp]] is a 3 chain structure with sequence from [https://en.wikipedia.org/wiki/Naja_kaouthia Naja kaouthia]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=3FRP OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=3FRP FirstGlance]. <br> |
| - | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=CA:CALCIUM+ION'>CA</scene></td></tr> | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.61Å</td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=CA:CALCIUM+ION'>CA</scene></td></tr> |
| + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=3frp FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=3frp OCA], [https://pdbe.org/3frp PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=3frp RCSB], [https://www.ebi.ac.uk/pdbsum/3frp PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=3frp ProSAT]</span></td></tr> | ||
</table> | </table> | ||
== Function == | == Function == | ||
| - | [ | + | [https://www.uniprot.org/uniprot/VCO3_NAJKA VCO3_NAJKA] Complement-activating protein in cobra venom. It is a structural and functional analog of complement component C3b, the activated form of C3. It binds factor B (CFB), which is subsequently cleaved by factor D (CFD) to form the bimolecular complex CVF/Bb. CVF/Bb is a C3/C5 convertase that cleaves both complement components C3 and C5. Structurally, it resembles the C3b degradation product C3c, which is not able to form a C3/C5 convertase. Unlike C3b/Bb, CVF/Bb is a stable complex and completely resistant to the actions of complement regulatory factors H (CFH) and I (CFI). Therefore, CVF continuously activates complement resulting in the depletion of complement activity. |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 37: | Line 38: | ||
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
[[Category: Naja kaouthia]] | [[Category: Naja kaouthia]] | ||
| - | [[Category: Krishnan | + | [[Category: Krishnan V]] |
| - | [[Category: Narayana | + | [[Category: Narayana SVL]] |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Revision as of 06:50, 6 September 2023
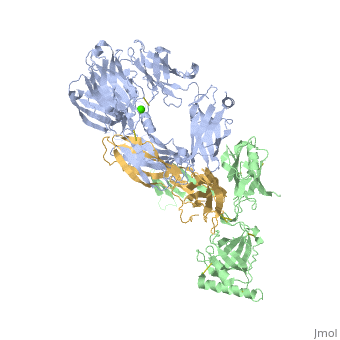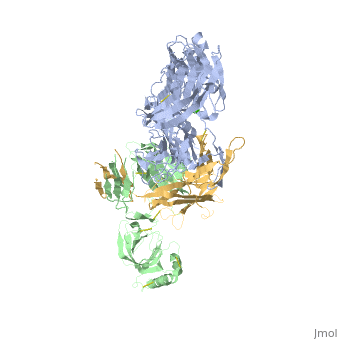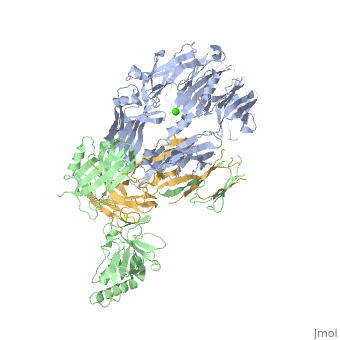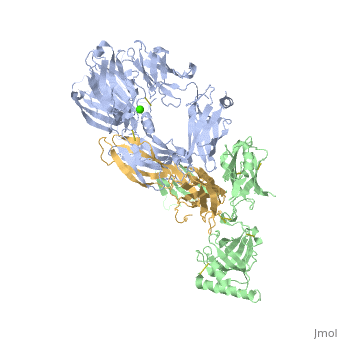
Crystal Structure of Cobra Venom Factor, a Co-factor for C3- and C5 convertase CVFBb
| |||||||||||