This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox GGC3
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
==Spike glycoprotein== | ==Spike glycoprotein== | ||
<StructureSection load='6VSB' size='340' side='right' caption='3D representation of the Spike glycoprotein' scene=''> | <StructureSection load='6VSB' size='340' side='right' caption='3D representation of the Spike glycoprotein' scene=''> | ||
| - | + | 3D structure representation of the Spike glycoprotein related to the SARS-CoV-2 <ref>DOI 10.1002/ijch.201300024</ref> <ref>PMID:21638687</ref>. | |
==Introduction== | ==Introduction== | ||
| Line 16: | Line 16: | ||
==Activation of S-protein== | ==Activation of S-protein== | ||
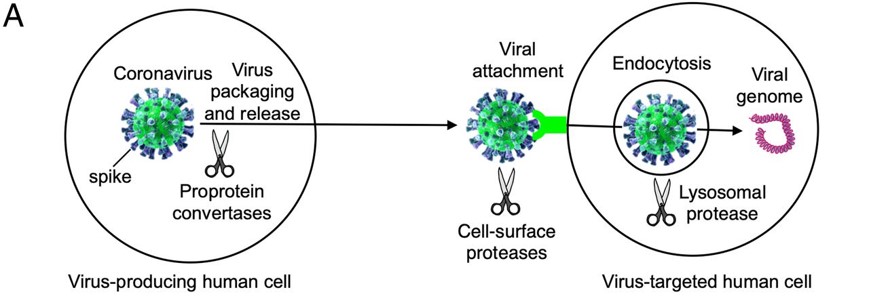
| - | + | Before the Spike protein can be activated , it has to be cleaved by the protease Furin protein ALA 668 <scene name='75/752266/Ala_668/1'>Furin Cleavage site</scene> . This 2D image below shows the schematic cleavage of the S-protein before and after it enters the host cell | |
[[Image:Cleavage.jpg]] <ref>Shang, J., Wan, Y., Luo, C., Ye, G., Geng, Q., Auerbach, A., & Li, F. (2020, May 26). Cell entry mechanisms of SARS-CoV-2. Retrieved November 14, 2020, from https://www.pnas.org/content/117/21/11727</ref> | [[Image:Cleavage.jpg]] <ref>Shang, J., Wan, Y., Luo, C., Ye, G., Geng, Q., Auerbach, A., & Li, F. (2020, May 26). Cell entry mechanisms of SARS-CoV-2. Retrieved November 14, 2020, from https://www.pnas.org/content/117/21/11727</ref> | ||
| Line 23: | Line 23: | ||
LYS 187 <scene name='75/752266/Lys_187/1'>The Receptor Binding Domain (RBD)</scene> | LYS 187 <scene name='75/752266/Lys_187/1'>The Receptor Binding Domain (RBD)</scene> | ||
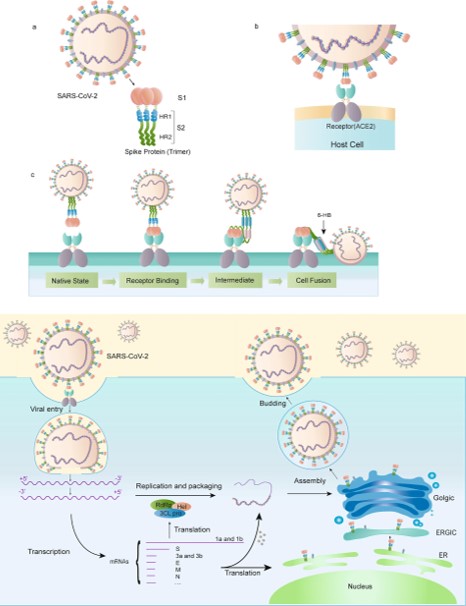
| - | [[Image:Picture3.jpg]]<ref> | + | [[Image:Picture3.jpg]]<ref>doi:10.1038/s41401-020-0485-4</ref> |
==Mutation== | ==Mutation== | ||
| - | + | The most studied mutation site of the S-protein is at residue 614 which encodes for the amino acid Aspartic acid (D) D614 <scene name='75/752266/Asp_614/1'>The Mutation site D614 </scene> and is normally changed to Glycine (G). And this form of mutation causes the enhancement of the viral transmission <ref>doi.org/10.1016/j.ijid.2020.10.033</ref>. | |
| Line 34: | Line 34: | ||
== Structural highlights == | == Structural highlights == | ||
| - | These are the structural 3D representations of the s-protein showing the two subunits , the binding regions to its receptor human ACE2 , Mutation site , RBD site and cleavage site respectively NTD-CT<scene name='75/752266/Ntd_-_ct/1'>S1 subunit is in the downstream of NTD and S2 subunit is in the upsteam of CT</scene> .Val367<scene name='75/752266/Val367/1'>Binding region val367</scene> . D614 <scene name='75/752266/Asp_614/1'>The Mutation site D614 </scene>. LYS 187 <scene name='75/752266/Lys_187/1'>The Receptor Binding Domain (RBD)</scene>.ALA 668 <scene name='75/752266/Ala_668/1'>Furin Cleavage site</scene> | + | These are the structural 3D representations of the s-protein showing the two subunits , the binding regions to its receptor human ACE2 , Mutation site , RBD site and cleavage site respectively. NTD-CT<scene name='75/752266/Ntd_-_ct/1'>S1 subunit is in the downstream of NTD and S2 subunit is in the upsteam of CT</scene> .Val367<scene name='75/752266/Val367/1'>Binding region val367</scene> . D614 <scene name='75/752266/Asp_614/1'>The Mutation site D614 </scene>. LYS 187 <scene name='75/752266/Lys_187/1'>The Receptor Binding Domain (RBD)</scene>.ALA 668 <scene name='75/752266/Ala_668/1'>Furin Cleavage site</scene> |
</StructureSection> | </StructureSection> | ||
Revision as of 18:05, 16 November 2020
Spike glycoprotein
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Huang, Y., Yang, C., Xu, X., Xu, W., & Liu, S. (2020). Structural and functional properties of SARS-CoV-2 spike protein: Potential antivirus drug development for COVID-19. Acta Pharmacologica Sinica, 41(9), 1141-1149. doi:10.1038/s41401-020-0485-4
- ↑ Mohammad, A., Alshawaf, E., Marafie, S. K., Abu-Farha, M., Abubaker, J., & Al-Mulla, F. (2020). Higher binding affinity of Furin to SARS-CoV-2 spike (S) protein D614G could be associated with higher SARS-CoV-2 infectivity. International journal of infectious diseases : IJID : official publication of the International Society for Infectious Diseases, S1201-9712(20)32237-2. Advance online publication. https://doi.org/10.1016/j.ijid.2020.10.033
- ↑ UniProt ConsortiumEuropean Bioinformatics InstituteProtein Information ResourceSIB Swiss Institute of Bioinformatics. (2020, October 07). Spike glycoprotein. Retrieved November 13, 2020, from https://www.uniprot.org/uniprot/P59594
- ↑ Shang, J., Wan, Y., Luo, C., Ye, G., Geng, Q., Auerbach, A., & Li, F. (2020, May 26). Cell entry mechanisms of SARS-CoV-2. Retrieved November 14, 2020, from https://www.pnas.org/content/117/21/11727
- ↑ Huang Y, Yang C, Xu XF, Xu W, Liu SW. Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacol Sin. 2020 Sep;41(9):1141-1149. doi: 10.1038/s41401-020-0485-4., Epub 2020 Aug 3. PMID:32747721 doi:http://dx.doi.org/10.1038/s41401-020-0485-4
- ↑ doi.org/10.1016/j.ijid.2020.10.033