|
|
| Line 3: |
Line 3: |
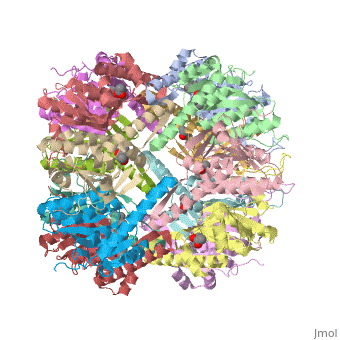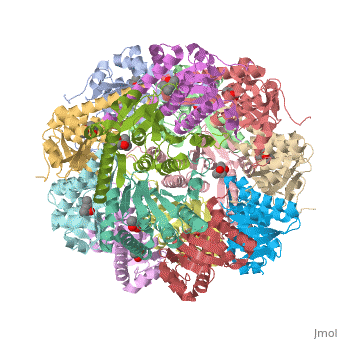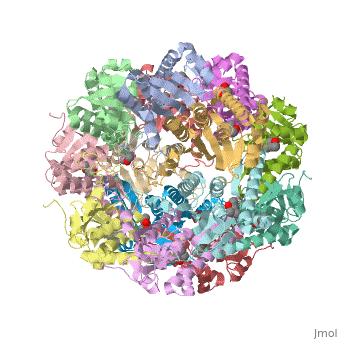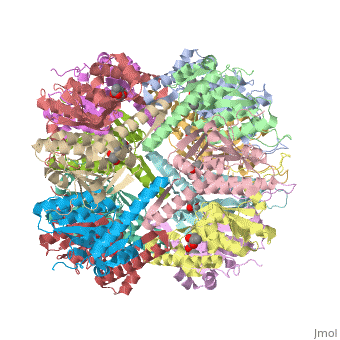
| | <StructureSection load='1yg6' size='340' side='right'caption='[[1yg6]], [[Resolution|resolution]] 1.90Å' scene=''> | | <StructureSection load='1yg6' size='340' side='right'caption='[[1yg6]], [[Resolution|resolution]] 1.90Å' scene=''> |
| | == Structural highlights == | | == Structural highlights == |
| - | <table><tr><td colspan='2'>[[1yg6]] is a 14 chain structure with sequence from [http://en.wikipedia.org/wiki/"bacillus_coli"_migula_1895 "bacillus coli" migula 1895]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1YG6 OCA]. For a <b>guided tour on the structure components</b> use [http://proteopedia.org/fgij/fg.htm?mol=1YG6 FirstGlance]. <br> | + | <table><tr><td colspan='2'>[[1yg6]] is a 14 chain structure with sequence from [https://en.wikipedia.org/wiki/Escherichia_coli Escherichia coli]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1YG6 OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1YG6 FirstGlance]. <br> |
| - | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=MPD:(4S)-2-METHYL-2,4-PENTANEDIOL'>MPD</scene></td></tr> | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 1.9Å</td></tr> |
| - | <tr id='related'><td class="sblockLbl"><b>[[Related_structure|Related:]]</b></td><td class="sblockDat"><div style='overflow: auto; max-height: 3em;'>[[1tyf|1tyf]]</div></td></tr>
| + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=MPD:(4S)-2-METHYL-2,4-PENTANEDIOL'>MPD</scene></td></tr> |
| - | <tr id='gene'><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">clpP, lopP ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=562 "Bacillus coli" Migula 1895])</td></tr> | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1yg6 FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1yg6 OCA], [https://pdbe.org/1yg6 PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1yg6 RCSB], [https://www.ebi.ac.uk/pdbsum/1yg6 PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1yg6 ProSAT]</span></td></tr> |
| - | <tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[http://en.wikipedia.org/wiki/Endopeptidase_Clp Endopeptidase Clp], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=3.4.21.92 3.4.21.92] </span></td></tr>
| + | |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://proteopedia.org/fgij/fg.htm?mol=1yg6 FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1yg6 OCA], [http://pdbe.org/1yg6 PDBe], [http://www.rcsb.org/pdb/explore.do?structureId=1yg6 RCSB], [http://www.ebi.ac.uk/pdbsum/1yg6 PDBsum], [http://prosat.h-its.org/prosat/prosatexe?pdbcode=1yg6 ProSAT]</span></td></tr> | + | |
| | </table> | | </table> |
| | == Function == | | == Function == |
| - | [[http://www.uniprot.org/uniprot/CLPP_ECOLI CLPP_ECOLI]] Cleaves peptides in various proteins in a process that requires ATP hydrolysis. Has a chymotrypsin-like activity. Plays a major role in the degradation of misfolded proteins. May play the role of a master protease which is attracted to different substrates by different specificity factors such as ClpA or ClpX. | + | [https://www.uniprot.org/uniprot/CLPP_ECOLI CLPP_ECOLI] Cleaves peptides in various proteins in a process that requires ATP hydrolysis. Has a chymotrypsin-like activity. Plays a major role in the degradation of misfolded proteins. May play the role of a master protease which is attracted to different substrates by different specificity factors such as ClpA or ClpX. |
| | == Evolutionary Conservation == | | == Evolutionary Conservation == |
| | [[Image:Consurf_key_small.gif|200px|right]] | | [[Image:Consurf_key_small.gif|200px|right]] |
| Line 34: |
Line 32: |
| | ==See Also== | | ==See Also== |
| | *[[Clp protease 3D structures|Clp protease 3D structures]] | | *[[Clp protease 3D structures|Clp protease 3D structures]] |
| - | *[[Molecular Playground/ClpP|Molecular Playground/ClpP]] | + | *[[Heat Shock Protein structures|Heat Shock Protein structures]] |
| | == References == | | == References == |
| | <references/> | | <references/> |
| | __TOC__ | | __TOC__ |
| | </StructureSection> | | </StructureSection> |
| - | [[Category: Bacillus coli migula 1895]] | + | [[Category: Escherichia coli]] |
| - | [[Category: Endopeptidase Clp]]
| + | |
| | [[Category: Large Structures]] | | [[Category: Large Structures]] |
| - | [[Category: Bewley, M C]] | + | [[Category: Bewley MC]] |
| - | [[Category: Flanagan, J M]] | + | [[Category: Flanagan JM]] |
| - | [[Category: Graziano, V]] | + | [[Category: Graziano V]] |
| - | [[Category: Griffin, K]] | + | [[Category: Griffin K]] |
| - | [[Category: Caseinolytic protease]]
| + | |
| - | [[Category: Endopeptidase clp]]
| + | |
| - | [[Category: Heat shock protein f21 5]]
| + | |
| - | [[Category: Hydrolase]]
| + | |
| - | [[Category: Protease ti]]
| + | |
| Structural highlights
Function
CLPP_ECOLI Cleaves peptides in various proteins in a process that requires ATP hydrolysis. Has a chymotrypsin-like activity. Plays a major role in the degradation of misfolded proteins. May play the role of a master protease which is attracted to different substrates by different specificity factors such as ClpA or ClpX.
Evolutionary Conservation
Check, as determined by ConSurfDB. You may read the explanation of the method and the full data available from ConSurf.
Publication Abstract from PubMed
ClpP is a self-compartmentalized proteolytic assembly comprised of two, stacked, heptameric rings that, when associated with its cognate hexameric ATPase (ClpA or ClpX), form the ClpAP and ClpXP ATP-dependent protease, respectively. The symmetry mismatch is an absolute feature of this large energy-dependent protease and also of the proteasome, which shares a similar barrel-shaped architecture, but how it is accommodated within the complex has yet to be understood, despite recent structural investigations, due in part to the conformational lability of the N-termini. We present the structures of Escherichia coli ClpP to 1.9A and an inactive variant that provide some clues for how this might be achieved. In the wild type protein, the highly conserved N-terminal 20 residues can be grouped into two major structural classes. In the first, a loop formed by residues 10-15 protrudes out of the central access channel extending approximately 12-15A from the surface of the oligomer resulting in the closing of the access channel observed in one ring. Similar loops are implied to be exclusively observed in human ClpP and a variant of ClpP from Streptococcus pneumoniae. In the other ring, a second class of loop is visible in the structure of wt ClpP from E. coli that forms closer to residue 16 and faces toward the interior of the molecule creating an open conformation of the access channel. In both classes, residues 18-20 provide a conserved interaction surface. In the inactive variant, a third class of N-terminal conformation is observed, which arises from a conformational change in the position of F17. We have performed a detailed functional analysis on each of the first 20 amino acid residues of ClpP. Residues that extend beyond the plane of the molecule (10-15) have a lesser effect on ATPase interaction than those lining the pore (1-7 and 16-20). Based upon our structure-function analysis, we present a model to explain the widely disparate effects of individual residues on ClpP-ATPase complex formation and also a possible functional reason for this mismatch.
The asymmetry in the mature amino-terminus of ClpP facilitates a local symmetry match in ClpAP and ClpXP complexes.,Bewley MC, Graziano V, Griffin K, Flanagan JM J Struct Biol. 2006 Feb;153(2):113-28. Epub 2005 Dec 1. PMID:16406682[1]
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.
See Also
References
- ↑ Bewley MC, Graziano V, Griffin K, Flanagan JM. The asymmetry in the mature amino-terminus of ClpP facilitates a local symmetry match in ClpAP and ClpXP complexes. J Struct Biol. 2006 Feb;153(2):113-28. Epub 2005 Dec 1. PMID:16406682 doi:10.1016/j.jsb.2005.09.011
|