This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Aminoacyl tRNA Synthetase
From Proteopedia
(Difference between revisions)
| Line 7: | Line 7: | ||
== 3D Structures of Aminoacyl tRNA synthetase == | == 3D Structures of Aminoacyl tRNA synthetase == | ||
[[Aminoacyl tRNA synthetase 3D structures]] | [[Aminoacyl tRNA synthetase 3D structures]] | ||
| + | |||
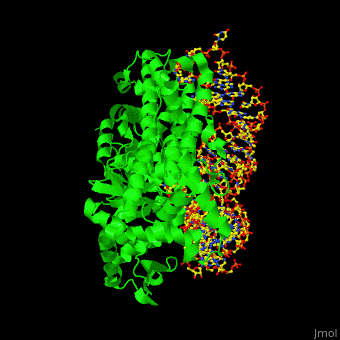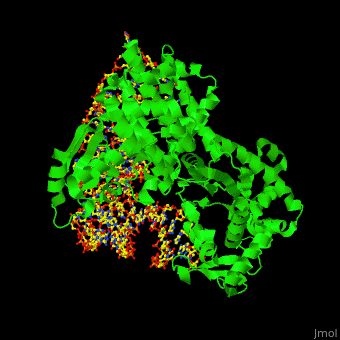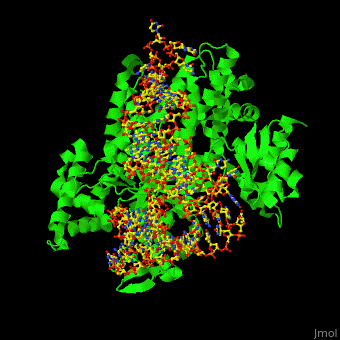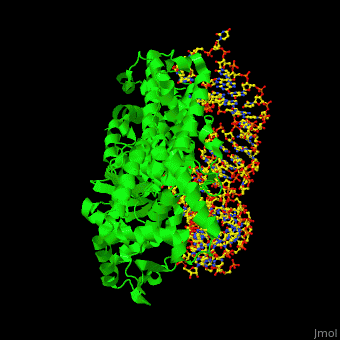
| + | One very interesting question in biology is how does an aminoacyl-tRNA synthetase recognize a particular tRNA and charge it with the correct amino acid? This is a challenging problem, since all tRNAs have the same general structure. Interestingly, different tRNA synthetases accomplish this goal in different ways. | ||
| + | |||
| + | The <scene name='44/444597/Protein_surface_interaction/1'>glutaminyl-tRNA synthetase</scene> (GlnRS) interacts with both the anticodon loop and the acceptor stem. Genetic and biochemical data indicate that GlnRS interacts with all three bases of the <scene name='44/444597/Anticodon_loop/1'>anticodon loop</scene>, which are unstacked and splay outward so they can bind in separate recognition pockets of GlnRS. The 3' end of tRNAgln plunges deeply into a protein pocket that also binds the enzymes ATP and glutamine substrates. | ||
</StructureSection> | </StructureSection> | ||
Revision as of 19:29, 21 September 2021
| |||||||||||
References
- ↑ Cavarelli J, Moras D. Recognition of tRNAs by aminoacyl-tRNA synthetases. FASEB J. 1993 Jan;7(1):79-86. PMID:8422978
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Joel L. Sussman, Ann Taylor