This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Leanne Price/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 23: | Line 23: | ||
===Active Site=== | ===Active Site=== | ||
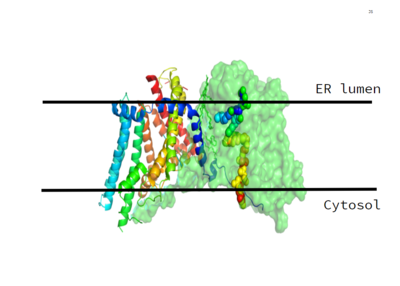
| - | The active site of DGAT is located within the membrane, with the catalytic histidine residue (<scene name='87/877601/His415/ | + | The active site of DGAT is located within the membrane, with the catalytic histidine residue (<scene name='87/877601/His415/2'>His415</scene>) buried inside the central cavity. This central cavity serves as the catalytic site. The acyl-acceptor lipid substrates access the active site through the lateral gate within the membrane. The active site also contains <scene name='87/877601/His415/1'>His415</scene> and several nearby polar residues (including Asn378, Gln437, and Gln465) whose side chains are oriented towards the cavity center. These residues interact and create a hydrophilic channel within the active site. The His415 residue is also likely involved in catalysis, making it increasingly significant. |
Revision as of 23:16, 17 March 2021
DGAT Human
| |||||||||||
References
- ↑ Wang L, Qian H, Nian Y, Han Y, Ren Z, Zhang H, Hu L, Prasad BVV, Laganowsky A, Yan N, Zhou M. Structure and mechanism of human diacylglycerol O-acyltransferase 1. Nature. 2020 May;581(7808):329-332. doi: 10.1038/s41586-020-2280-2. Epub 2020 May, 13. PMID:32433610 doi:http://dx.doi.org/10.1038/s41586-020-2280-2
- ↑ Sui X, Wang K, Gluchowski NL, Elliott SD, Liao M, Walther TC, Farese RV Jr. Structure and catalytic mechanism of a human triacylglycerol-synthesis enzyme. Nature. 2020 May;581(7808):323-328. doi: 10.1038/s41586-020-2289-6. Epub 2020 May, 13. PMID:32433611 doi:http://dx.doi.org/10.1038/s41586-020-2289-6
Student Contributors
- Justin Smith
- Eloi Bigirimana
- Leanne Price