This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Sarah Maarouf/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 19: | Line 19: | ||
=== GPIHBP1 === | === GPIHBP1 === | ||
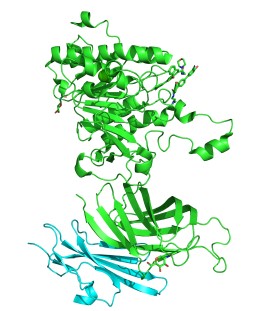
| - | <scene name='87/877554/Lpl-gpihbp1/ | + | GPIHBP1 is a membrane bound protein that binds to the C-terminal domain of LPL. It contains specific features, the LU domain and the acidic domain, that contribute to its binding affinity. GPIHBP1’s LU domain is 75 residues in length and adopts a <scene name='87/877554/Lpl-gpihbp1/5'>three-fingered fold</scene> that is stabilized by <scene name='87/877554/Disulfide_bonds_gpihbp1/2'>five disulfide bonds</scene>. The LPL-GPIHBP1 binding interface involves the <scene name='87/877554/Lpl-gpihbp1/5'>three-fingered fold of GPIHBP1</scene> and depends largely on hydrophobic interactions between the two subunits. It contains a highly acidic and disordered N-terminal domain with 21 of the 26 total residues being aspartates or glutamates which were not visible in the electron density map. This indicates that the acidic domain of GPIHBP1 likely exhibits conformational flexibility. |
| - | + | ||
| - | <scene name='87/877554/ | + | |
== Function == | == Function == | ||
Revision as of 14:49, 3 April 2021
H. sapiens Lipoprotein Lipase in complex with GPIHBP1
| |||||||||||
New text we are referencing [3]
References
- ↑ Birrane G, Beigneux AP, Dwyer B, Strack-Logue B, Kristensen KK, Francone OL, Fong LG, Mertens HDT, Pan CQ, Ploug M, Young SG, Meiyappan M. Structure of the lipoprotein lipase-GPIHBP1 complex that mediates plasma triglyceride hydrolysis. Proc Natl Acad Sci U S A. 2018 Dec 17. pii: 1817984116. doi:, 10.1073/pnas.1817984116. PMID:30559189 doi:http://dx.doi.org/10.1073/pnas.1817984116
- ↑ Arora R, Nimonkar AV, Baird D, Wang C, Chiu CH, Horton PA, Hanrahan S, Cubbon R, Weldon S, Tschantz WR, Mueller S, Brunner R, Lehr P, Meier P, Ottl J, Voznesensky A, Pandey P, Smith TM, Stojanovic A, Flyer A, Benson TE, Romanowski MJ, Trauger JW. Structure of lipoprotein lipase in complex with GPIHBP1. Proc Natl Acad Sci U S A. 2019 May 21;116(21):10360-10365. doi:, 10.1073/pnas.1820171116. Epub 2019 May 9. PMID:31072929 doi:http://dx.doi.org/10.1073/pnas.1820171116
- ↑ 3.0 3.1 Ransey E, Paredes E, Dey SK, Das SR, Heroux A, Macbeth MR. Crystal structure of the Entamoeba histolytica RNA lariat debranching enzyme EhDbr1 reveals a catalytic Zn(2+) /Mn(2+) heterobinucleation. FEBS Lett. 2017 Jul;591(13):2003-2010. doi: 10.1002/1873-3468.12677. Epub 2017, Jun 14. PMID:28504306 doi:http://dx.doi.org/10.1002/1873-3468.12677
Student Contributors
- Aniyah Coles
- Sarah Maarouf
- Audrey Marjamaa