This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Leanne Price/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 31: | Line 31: | ||
===Active Site=== | ===Active Site=== | ||
| - | The active site of DGAT is located within the membrane, with the catalytic histidine residue (<scene name='87/877601/His415/5'>His415</scene>-represented in white) buried inside the central cavity. This central cavity serves as the catalytic site. The acyl-acceptor lipid substrates access the active site through the lateral gate within the membrane. The active site also contains <scene name='87/877601/His415_tunnel/2'>His415</scene> and several nearby <scene name='87/877601/His415_and_polar_residues/3'>polar residues</scene> (including Asn378, Gln437, and Gln465) whose side chains are oriented towards the cavity center. These residues interact and create a hydrophilic channel within the active site. The His415 residue is also likely involved in catalysis, making it increasingly significant. In face, single mutations of His415 and Asn378 terminated DGAT activity. This suggests that the central cavity of DGAT within the membrane is the catalytic site. | + | The active site of DGAT is located within the membrane, with the catalytic histidine residue (<scene name='87/877601/His415/5'>His415</scene>-represented in white) buried inside the central cavity. This central cavity serves as the catalytic site. The acyl-acceptor lipid substrates access the active site through the lateral gate within the membrane. The active site also contains <scene name='87/877601/His415_tunnel/2'>His415</scene> and several nearby <scene name='87/877601/His415_and_polar_residues/3'>polar residues</scene> (including Asn378, Gln437, and Gln465) whose side chains are oriented towards the cavity center. These residues interact and create a hydrophilic channel within the active site. The His415 residue is also likely involved in catalysis, making it increasingly significant. In face, single mutations of His415 and Asn378 terminated DGAT activity. This suggests that the central cavity of DGAT within the membrane is the catalytic site. <ref name="Sui" /ref> |
| + | |||
==Mechanism== | ==Mechanism== | ||
| Line 39: | Line 40: | ||
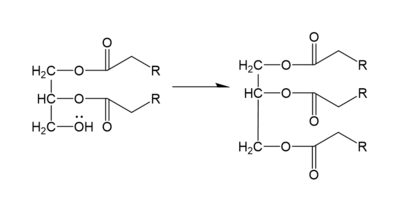
In the DGAT mechanism, the diglyceride serves as the nucleophile. While the acyl group of the CoA enzyme serves as the electrophile. The lone pair on the last hydroxyl group present on the glycerol of the diglyceride attacks the thioester bond of the acyl-CoA enzyme. This attack breaks the sulfur-carbon bond, a weak bond that is easily breakable. This allows the acyl group of the acyl-CoA enzyme to attach to the diglyceride, creating a triglyceride. While the CoA group then serves as the leaving group. | In the DGAT mechanism, the diglyceride serves as the nucleophile. While the acyl group of the CoA enzyme serves as the electrophile. The lone pair on the last hydroxyl group present on the glycerol of the diglyceride attacks the thioester bond of the acyl-CoA enzyme. This attack breaks the sulfur-carbon bond, a weak bond that is easily breakable. This allows the acyl group of the acyl-CoA enzyme to attach to the diglyceride, creating a triglyceride. While the CoA group then serves as the leaving group. | ||
| - | The cleavage site for oleoyl-CoA is within a short distance of a lipid acceptor. This revelation was made after a strong, lipid-like density in the central cavity in the cryo-EM data. Hydrophobic residues line this region and form a channel surrounding the lipid-like density. The channel itself has a bent, hydrophobic pathway that allows the binding of hydrophobic molecules. The bent architecture of this tunnel is likely how DGAT distinguishes acyl acceptors from other molecules, such as cholesterol. | + | The cleavage site for oleoyl-CoA is within a short distance of a lipid acceptor. This revelation was made after a strong, lipid-like density in the central cavity in the cryo-EM data. Hydrophobic residues line this region and form a channel surrounding the lipid-like density. The channel itself has a bent, hydrophobic pathway that allows the binding of hydrophobic molecules. The bent architecture of this tunnel is likely how DGAT distinguishes acyl acceptors from other molecules, such as cholesterol. <ref name="Sui" /ref> |
| + | |||
===Product Release=== | ===Product Release=== | ||
| - | As previously mentioned, the Acyl-CoA molecule serves as the leaving group in the DGAT mechanism. This acyl-CoA molecule occupies the cytosolic tunnel, which has a bent architecture. The CoA moiety is at the cytosolic face, while the acyl chain extends through the center towards the endoplasmic reticulum lumen. The distal end of the acyl chain oleoyl-CoA interacts with DGAT deep within the hydrophobic channel, which suggests that the binding of longer acyl chains help accurately position the acyl-donor substrate for the reaction. As the acyl-CoA binds to DGAT, small conformational changes are seen in the active site region, specifically the His415 residue flips towards the endoplasmic reticulum-luminal side when acyl-CoA binds. This conformational change allows a new hydrogen bond to form and positions His415 near the thioester bond of the acyl-CoA. Therefore, the binding of acyl-CoA binding to DGAT results in small, but important, conformational changes in the active site that likely prime the enzyme for catalysis. | + | As previously mentioned, the Acyl-CoA molecule serves as the leaving group in the DGAT mechanism. This acyl-CoA molecule occupies the cytosolic tunnel, which has a bent architecture. The CoA moiety is at the cytosolic face, while the acyl chain extends through the center towards the endoplasmic reticulum lumen. The distal end of the acyl chain oleoyl-CoA interacts with DGAT deep within the hydrophobic channel, which suggests that the binding of longer acyl chains help accurately position the acyl-donor substrate for the reaction. As the acyl-CoA binds to DGAT, small conformational changes are seen in the active site region, specifically the His415 residue flips towards the endoplasmic reticulum-luminal side when acyl-CoA binds. This conformational change allows a new hydrogen bond to form and positions His415 near the thioester bond of the acyl-CoA. Therefore, the binding of acyl-CoA binding to DGAT results in small, but important, conformational changes in the active site that likely prime the enzyme for catalysis. <ref name="Sui" /ref> |
| + | |||
==Mutations== | ==Mutations== | ||
Revision as of 19:29, 13 April 2021
Diacylglycerol Acyltransferase
| |||||||||||
References
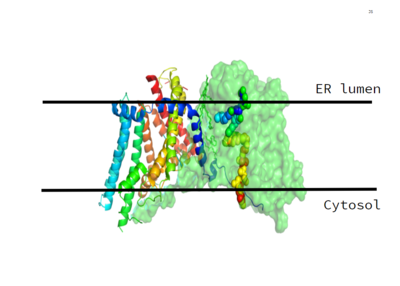
- ↑ 1.0 1.1 1.2 DGAT was originally discovered by its homology to Acyl-CoA cholesterol acyltransferases (ACAT) 1 and 2. The structure, catalytic mechanism of diacylglycerol acyltransferase, and how DGAT interacts with CoA was discovered using a Cryo-EM. The Cryo-EM map revealed that DGAT forms a dimer, with each subunit containing nine transmembrane helices. The N and C terminals of each helix are located on the cytosolic and luminal sides of the endoplasmic reticulum membrane respectively (Figure 1). <ref>PMID:32433610</li></ol></ref>
Student Contributors
- Justin Smith
- Eloi Bigirimana
- Leanne Price