This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Kaitlyn Roberts/Sandbox 2
From Proteopedia
(Difference between revisions)
| Line 9: | Line 9: | ||
== Structure == | == Structure == | ||
=== Tertiary Structure === | === Tertiary Structure === | ||
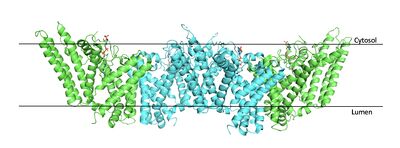
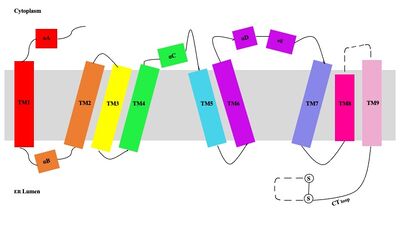
| - | [[Image:Tetramerlabels.jpeg|400 px|right|thumb|'''Figure 2. Tetramer unit of SOAT shown in position within the membrane.''' The dimer units are identical, as indicated by the corresponding green and blue regions. [http://www.rcsb.org/structure/6P2P PBD 6P2P]]] The biological assembly of SOAT is a <scene name='87/877559/Tetramer/11'>tetramer</scene> or a <scene name='87/877559/Tetramer/10'>dimer of dimers</scene>. Functionally, the <scene name='87/877559/Dimer/3'>dimer</scene> units of SOAT are identical and | + | [[Image:Tetramerlabels.jpeg|400 px|right|thumb|'''Figure 2. Tetramer unit of SOAT shown in position within the membrane.''' The dimer units are identical, as indicated by the corresponding green and blue regions. [http://www.rcsb.org/structure/6P2P PBD 6P2P]]] The biological assembly of SOAT is a <scene name='87/877559/Tetramer/11'>tetramer</scene> or a <scene name='87/877559/Tetramer/10'>dimer of dimers</scene>. Functionally, the <scene name='87/877559/Dimer/3'>dimer</scene> units of SOAT are identical and are stabilized by hydrophobic [https://en.wikipedia.org/wiki/Van_der_Waals_force van der Waals interactions] between residues at the <scene name='87/877559/Dimer_interface/3'>dimer interface</scene>. Mutating these residues inhibits enzyme activity, suggesting that the dimer unit of SOAT is critical for enzyme function.<ref name="Guan" /> Each dimer consists of two identical <scene name='87/877559/Monomer/5'>monomer</scene> units, individually made up of nine <scene name='87/877559/Helices_1-9/4'>transmembrane helices</scene> labeled TM1 through TM9. [[Image:Helicesdiagram1.jpeg|400 px|right|thumb|'''Figure 3.''' Labeled helices of SOAT within the membrane]] The van der Waals interactions at the dimer interface stabilize the dimer between the TM1 helice of one monomer unit and the TM6 [https://en.wikipedia.org/wiki/Lumen_(anatomy) lumenal] segment and TM9 [https://en.wikipedia.org/wiki/Cytosol cytosolic] segment of the other monomer unit. Essential helices (TM1, TM5, TM6, and TM9) from the two monomers form the entrance tunnels and catalytic active site.<ref name="Qian">PMID:32433614</ref> |
=== Tunnel System === | === Tunnel System === | ||
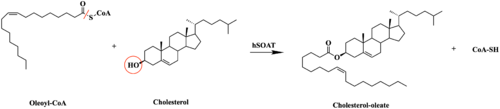
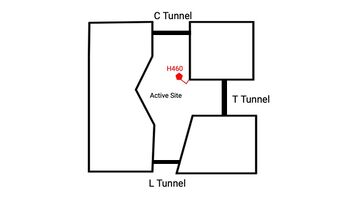
| - | + | An important structural element of SOAT is the tunnel system through which substrates enter and exit. [[Image:Tunnels2.jpg|350 px|right|thumb|'''Figure 4. 2D layout of the SOAT tunnel system.''' The orientation of the tunnels shows the C tunnel opening to the cytosol and the L tunnel opening to the lumen. The T tunnel opens into the membrane, but is not quite the 90 degree shown in the 2D image.]] There are 3 main tunnels in each monomer: the cytosolic (C) tunnel opening to the cytosol, the transmembrane(T) tunnel opening to the membrane, and the lumenal (L) tunnel opens to the lumen. <ref name="Qian" /> The C tunnel opens to the cytosol of the cell and is the entrance site for the Acyl CoA substrate into the active site. Residues <scene name='87/877559/C_tunnel_and_measurements/6'>N415, Y433, and K445</scene> are involved in hydrogen bonding interactions with polar atoms on CoenzymeA to help stabilize the substrate within the binding pocket. It is important to note that the N415 residue in the PDB file provided is likely incorrect in orientation. We believe it should be flipped so that the hydrogens associated with the nitrogen atom are performing the stabilization shown. Surface representations of SOAT indicate that there are 2 alpha helices that block the entrance to the C tunnel, therefore a conformational change needs to occur to move the 2 helices so the substrate can enter the tunnel. The T tunnel opens into the membrane and is where cholesterol enters to have access to the active site. The two substrates are catalyzed by the H460 in the active site to form the cholesteryl ester. The products then leave via different pathways. The CoA-SH in the C tunnel leaves via that tunnel and is released back into the cytosol. The cholesteryl ester then leaves via either the T tunnel into the membrane or through the L tunnel into the lumen of the cell. <ref name="Qian" /> | |
=== Active Site === | === Active Site === | ||
Revision as of 21:36, 26 April 2021
Human Sterol O-acyltransferase
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 Guan C, Niu Y, Chen SC, Kang Y, Wu JX, Nishi K, Chang CCY, Chang TY, Luo T, Chen L. Structural insights into the inhibition mechanism of human sterol O-acyltransferase 1 by a competitive inhibitor. Nat Commun. 2020 May 18;11(1):2478. doi: 10.1038/s41467-020-16288-4. PMID:32424158 doi:http://dx.doi.org/10.1038/s41467-020-16288-4
- ↑ 2.0 2.1 2.2 2.3 Qian H, Zhao X, Yan R, Yao X, Gao S, Sun X, Du X, Yang H, Wong CCL, Yan N. Structural basis for catalysis and substrate specificity of human ACAT1. Nature. 2020 May;581(7808):333-338. doi: 10.1038/s41586-020-2290-0. Epub 2020 May, 13. PMID:32433614 doi:http://dx.doi.org/10.1038/s41586-020-2290-0
- ↑ Das A, Davis MA, Rudel LL. Identification of putative active site residues of ACAT enzymes. J Lipid Res. 2008 Aug;49(8):1770-81. doi: 10.1194/jlr.M800131-JLR200. Epub 2008, May 13. PMID:18480028 doi:http://dx.doi.org/10.1194/jlr.M800131-JLR200
- ↑ Guo ZY, Lin S, Heinen JA, Chang CC, Chang TY. The active site His-460 of human acyl-coenzyme A:cholesterol acyltransferase 1 resides in a hitherto undisclosed transmembrane domain. J Biol Chem. 2005 Nov 11;280(45):37814-26. doi: 10.1074/jbc.M508384200. Epub 2005, Sep 8. PMID:16154994 doi:http://dx.doi.org/10.1074/jbc.M508384200
- ↑ 5.0 5.1 Bhattacharyya R, Kovacs DM. ACAT inhibition and amyloid beta reduction. Biochim Biophys Acta. 2010 Aug;1801(8):960-5. doi: 10.1016/j.bbalip.2010.04.003. , Epub 2010 Apr 14. PMID:20398792 doi:http://dx.doi.org/10.1016/j.bbalip.2010.04.003
- ↑ 6.0 6.1 Huttunen HJ, Kovacs DM. ACAT as a drug target for Alzheimer's disease. Neurodegener Dis. 2008;5(3-4):212-4. doi: 10.1159/000113705. Epub 2008 Mar 6. PMID:18322393 doi:http://dx.doi.org/10.1159/000113705
- ↑ Chang C, Dong R, Miyazaki A, Sakashita N, Zhang Y, Liu J, Guo M, Li BL, Chang TY. Human acyl-CoA:cholesterol acyltransferase (ACAT) and its potential as a target for pharmaceutical intervention against atherosclerosis. Acta Biochim Biophys Sin (Shanghai). 2006 Mar;38(3):151-6. doi:, 10.1111/j.1745-7270.2006.00154.x. PMID:16518538 doi:http://dx.doi.org/10.1111/j.1745-7270.2006.00154.x
- ↑ Ayyagari VN, Wang X, Diaz-Sylvester PL, Groesch K, Brard L. Assessment of acyl-CoA cholesterol acyltransferase (ACAT-1) role in ovarian cancer progression-An in vitro study. PLoS One. 2020 Jan 24;15(1):e0228024. doi: 10.1371/journal.pone.0228024., eCollection 2020. PMID:31978092 doi:http://dx.doi.org/10.1371/journal.pone.0228024
Student Contributors
- Kylie Pfeifer
- Stephanie Pellegrino
- Kaitlyn Roberts