This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1715
From Proteopedia
(Difference between revisions)
| Line 31: | Line 31: | ||
|} | |} | ||
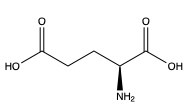
| - | [[Image:Screen Shot 2022-03-28 at 8.44.00 AM.png|300 px|right|thumb|Figure | + | [[Image:Screen Shot 2022-03-28 at 8.44.00 AM.png|300 px|right|thumb|Figure 1. Structure of glutamate. ]]In order to activate the mGlu transformation, glutamate acts as the protein’s main agonist. Glutamate is an acidic, polar amino acid. (picture) This agonist binds to the extracellular portion of the protein allowing the homodimer to change conformationally. This change allows for a signaling cascade within the cell that can ultimately lead to the modification of other proteins and a difference in the synapse’s excitability. However, in mGlu, the binding affinity of glutamate is determined by either a positive (PAM) or negative (NAM) allosteric modulator. |
== Structural Highlights == | == Structural Highlights == | ||
mGlu receptors are dimeric proteins consisting of an <scene name='90/904320/Inactive_mglu2/1'>alpha and beta chain</scene>. While a heterodimer of different mGlu subtypes can form, only homodimeric receptors can become active. Both the alpha and beta chains are comprised of <scene name='90/904320/Mglu2_domains/2'>3 domains</scene>: the venus fly trap (VFT), cysteine rich domain (CRD), and the transmembrane domain (TMD). | mGlu receptors are dimeric proteins consisting of an <scene name='90/904320/Inactive_mglu2/1'>alpha and beta chain</scene>. While a heterodimer of different mGlu subtypes can form, only homodimeric receptors can become active. Both the alpha and beta chains are comprised of <scene name='90/904320/Mglu2_domains/2'>3 domains</scene>: the venus fly trap (VFT), cysteine rich domain (CRD), and the transmembrane domain (TMD). | ||
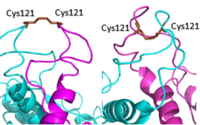
| - | [[Image: Active vs inactive vft.png|200px|left|thumb|Figure | + | [[Image: Active vs inactive vft.png|200px|left|thumb|Figure 2. Cysteine 121 positioning in the inactive (left) vs the active (right) mGlu conformation ]] |
==== Domains ==== | ==== Domains ==== | ||
<scene name='90/904320/Vft/2'>VFT</scene>: The extracellular location in which the two glutamate agonists bind is known as the VFT. This domain includes a disulfide bond which is shifted down in the inactive form and undergoes an upward movement upon glutamate binding which stabilizes the active site (Figure 1). | <scene name='90/904320/Vft/2'>VFT</scene>: The extracellular location in which the two glutamate agonists bind is known as the VFT. This domain includes a disulfide bond which is shifted down in the inactive form and undergoes an upward movement upon glutamate binding which stabilizes the active site (Figure 1). | ||
| - | <scene name='90/904320/Crd/ | + | <scene name='90/904320/Crd/5'>CRD</scene>: The portion of the protomer that connects the VFT with the TMD is known as the CRD. As the linking segment of the protein, is critical in transmitting the conformational change caused by the binding of glutamate to the TMD. The change resulting from the binding of glutamate in the VFT brings the cysteine-rich domain together to alter the configuration of the TMD through its interaction with the extracellular loop 2 (ECL2). This process is mediated by the hydrophobic effect due to the nature of the amino acids at the apex of the CRD. |
'''TMD''': The TMD consists of seven transmembrane helices that are responsible for G-protein interactions. In the inactive form, the asymmetric conformation of the helices is mediated by the hydrophobicity of helix 3 and 4. Along with the interaction of the CRD with the ECL2 of the TMD, an allosteric modulator must bind within the transmembrane helices to allow for the confirmation of the helices to be altered. This conformation allows for a dimer interface along helix 6 (citation article 2) . The stabilization of this conformation allows for G protein coupling with ICL2, ICL3, TM Helix 3 and the C terminus. | '''TMD''': The TMD consists of seven transmembrane helices that are responsible for G-protein interactions. In the inactive form, the asymmetric conformation of the helices is mediated by the hydrophobicity of helix 3 and 4. Along with the interaction of the CRD with the ECL2 of the TMD, an allosteric modulator must bind within the transmembrane helices to allow for the confirmation of the helices to be altered. This conformation allows for a dimer interface along helix 6 (citation article 2) . The stabilization of this conformation allows for G protein coupling with ICL2, ICL3, TM Helix 3 and the C terminus. | ||
Revision as of 22:30, 28 March 2022
Metabotropic Glutamate Receptor
| |||||||||||
References
- ↑ 1.0 1.1 Ransey E, Paredes E, Dey SK, Das SR, Heroux A, Macbeth MR. Crystal structure of the Entamoeba histolytica RNA lariat debranching enzyme EhDbr1 reveals a catalytic Zn(2+) /Mn(2+) heterobinucleation. FEBS Lett. 2017 Jul;591(13):2003-2010. doi: 10.1002/1873-3468.12677. Epub 2017, Jun 14. PMID:28504306 doi:http://dx.doi.org/10.1002/1873-3468.12677
- ↑ 2.0 2.1 Seven AB, Barros-Alvarez X, de Lapeyriere M, Papasergi-Scott MM, Robertson MJ, Zhang C, Nwokonko RM, Gao Y, Meyerowitz JG, Rocher JP, Schelshorn D, Kobilka BK, Mathiesen JM, Skiniotis G. G-protein activation by a metabotropic glutamate receptor. Nature. 2021 Jun 30. pii: 10.1038/s41586-021-03680-3. doi:, 10.1038/s41586-021-03680-3. PMID:34194039 doi:http://dx.doi.org/10.1038/s41586-021-03680-3
Student Contributors
- Courtney Vennekotter
- Cade Chezem