This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Kiera Malone/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 26: | Line 26: | ||
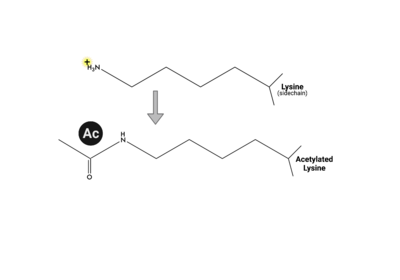
Epigenetics is the study of the molecules or mechanisms that cause changes in gene expression, without any modification to the underlying DNA sequence. DNA methylation, long non-coding RNAs, and post-translational modifications (PTMs) are all areas of epigenetic study (waddington). | Epigenetics is the study of the molecules or mechanisms that cause changes in gene expression, without any modification to the underlying DNA sequence. DNA methylation, long non-coding RNAs, and post-translational modifications (PTMs) are all areas of epigenetic study (waddington). | ||
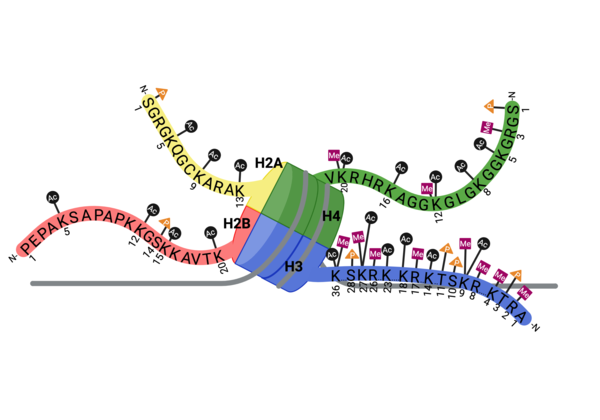
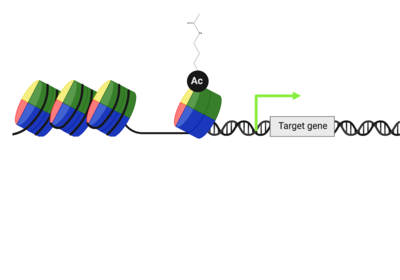
| - | PTMs can occur on the histone proteins that compose the <scene name='90/909366/Ncp/1'>nucleosome core particle</scene> (NCP). The NCP is the fundamental unit of chromatin and impacts gene accessibility to transcriptional regulators. Histones are positively charged and work to compact the negatively charged DNA around an octamer of histone proteins. Additionally, there are unstructured extensions at the N-terminus of histone proteins, termed <scene name='90/909366/Ncp/2'>tails</scene>, which are also positively charged. This compact histone octamer wrapped with DNA becomes the nucleosome core particle (NCP), the fundamental unit of chromatin, which poses a barrier to transcription (strahl)(luger). | + | PTMs can occur on the histone proteins that compose the <scene name='90/909366/Ncp/1'>nucleosome core particle</scene> (NCP). The NCP is the fundamental unit of chromatin and impacts gene accessibility to transcriptional regulators. Histones are positively charged and work to compact the negatively charged DNA around an octamer of histone proteins. Multiple NCPs compact DNA to form higher order chromatin structure, all the way to the level of the chromosome(CAVALLI). Additionally, there are unstructured extensions at the N-terminus of histone proteins, termed <scene name='90/909366/Ncp/2'>tails</scene>, which are also positively charged. This compact histone octamer wrapped with DNA becomes the nucleosome core particle (NCP), the fundamental unit of chromatin, which poses a barrier to transcription (strahl)(luger). |
[[Image:Postptms.png|center|600px]] | [[Image:Postptms.png|center|600px]] | ||
Revision as of 18:25, 30 April 2022
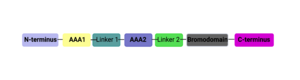
The ATPase Family, AAA Domain-Containing Protein 2B (ATAD2B)
| |||||||||||
References
- ↑ Leachman NT, Brellier F, Ferralli J, Chiquet-Ehrismann R, Tucker RP. ATAD2B is a phylogenetically conserved nuclear protein expressed during neuronal differentiation and tumorigenesis. Dev Growth Differ. 2010 Dec;52(9):747-55. doi: 10.1111/j.1440-169X.2010.01211.x. PMID:21158754 doi:http://dx.doi.org/10.1111/j.1440-169X.2010.01211.x
- ↑ Caron C, Lestrat C, Marsal S, Escoffier E, Curtet S, Virolle V, Barbry P, Debernardi A, Brambilla C, Brambilla E, Rousseaux S, Khochbin S. Functional characterization of ATAD2 as a new cancer/testis factor and a predictor of poor prognosis in breast and lung cancers. Oncogene. 2010 Sep 16;29(37):5171-81. doi: 10.1038/onc.2010.259. Epub 2010 Jun, 28. PMID:20581866 doi:http://dx.doi.org/10.1038/onc.2010.259
- ↑ Kalashnikova EV, Revenko AS, Gemo AT, Andrews NP, Tepper CG, Zou JX, Cardiff RD, Borowsky AD, Chen HW. ANCCA/ATAD2 overexpression identifies breast cancer patients with poor prognosis, acting to drive proliferation and survival of triple-negative cells through control of B-Myb and EZH2. Cancer Res. 2010 Nov 15;70(22):9402-12. doi: 10.1158/0008-5472.CAN-10-1199. Epub , 2010 Sep 23. PMID:20864510 doi:http://dx.doi.org/10.1158/0008-5472.CAN-10-1199
- ↑ Filippakopoulos P, Picaud S, Mangos M, Keates T, Lambert JP, Barsyte-Lovejoy D, Felletar I, Volkmer R, Muller S, Pawson T, Gingras AC, Arrowsmith CH, Knapp S. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell. 2012 Mar 30;149(1):214-31. PMID:22464331 doi:10.1016/j.cell.2012.02.013
- ↑ Evans CM, Phillips M, Malone KL, Tonelli M, Cornilescu G, Cornilescu C, Holton SJ, Gorjanacz M, Wang L, Carlson S, Gay JC, Nix JC, Demeler B, Markley JL, Glass KC. Coordination of Di-Acetylated Histone Ligands by the ATAD2 Bromodomain. Int J Mol Sci. 2021 Aug 24;22(17). pii: ijms22179128. doi: 10.3390/ijms22179128. PMID:34502039 doi:http://dx.doi.org/10.3390/ijms22179128
- ↑ Lloyd JT, McLaughlin K, Lubula MY, Gay JC, Dest A, Gao C, Phillips M, Tonelli M, Cornilescu G, Marunde MR, Evans CM, Boyson SP, Carlson S, Keogh MC, Markley JL, Frietze S, Glass KC. Structural Insights into the Recognition of Mono- and Diacetylated Histones by the ATAD2B Bromodomain. J Med Chem. 2020 Oct 21. doi: 10.1021/acs.jmedchem.0c01178. PMID:33084328 doi:http://dx.doi.org/10.1021/acs.jmedchem.0c01178
- ↑ Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Zidek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera-Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D. Highly accurate protein structure prediction with AlphaFold. Nature. 2021 Jul 15. pii: 10.1038/s41586-021-03819-2. doi:, 10.1038/s41586-021-03819-2. PMID:34265844 doi:http://dx.doi.org/10.1038/s41586-021-03819-2
- ↑ Varadi M, Anyango S, Deshpande M, Nair S, Natassia C, Yordanova G, Yuan D, Stroe O, Wood G, Laydon A, Zidek A, Green T, Tunyasuvunakool K, Petersen S, Jumper J, Clancy E, Green R, Vora A, Lutfi M, Figurnov M, Cowie A, Hobbs N, Kohli P, Kleywegt G, Birney E, Hassabis D, Velankar S. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 2022 Jan 7;50(D1):D439-D444. doi: 10.1093/nar/gkab1061. PMID:34791371 doi:http://dx.doi.org/10.1093/nar/gkab1061
- ↑ Filippakopoulos P, Picaud S, Mangos M, Keates T, Lambert JP, Barsyte-Lovejoy D, Felletar I, Volkmer R, Muller S, Pawson T, Gingras AC, Arrowsmith CH, Knapp S. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell. 2012 Mar 30;149(1):214-31. PMID:22464331 doi:10.1016/j.cell.2012.02.013