This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
4g9p
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
== Structural highlights == | == Structural highlights == | ||
<table><tr><td colspan='2'>[[4g9p]] is a 1 chain structure with sequence from [https://en.wikipedia.org/wiki/Thermus_thermophilus_HB27 Thermus thermophilus HB27]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=4G9P OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=4G9P FirstGlance]. <br> | <table><tr><td colspan='2'>[[4g9p]] is a 1 chain structure with sequence from [https://en.wikipedia.org/wiki/Thermus_thermophilus_HB27 Thermus thermophilus HB27]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=4G9P OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=4G9P FirstGlance]. <br> | ||
| - | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=CDI:2C-METHYL-D-ERYTHRITOL+2,4-CYCLODIPHOSPHATE'>CDI</scene>, <scene name='pdbligand=CL:CHLORIDE+ION'>CL</scene>, <scene name='pdbligand=K:POTASSIUM+ION'>K</scene>, <scene name='pdbligand=MES:2-(N-MORPHOLINO)-ETHANESULFONIC+ACID'>MES</scene>, <scene name='pdbligand=SF4:IRON/SULFUR+CLUSTER'>SF4</scene></td></tr> | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 1.55Å</td></tr> |
| + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=CDI:2C-METHYL-D-ERYTHRITOL+2,4-CYCLODIPHOSPHATE'>CDI</scene>, <scene name='pdbligand=CL:CHLORIDE+ION'>CL</scene>, <scene name='pdbligand=K:POTASSIUM+ION'>K</scene>, <scene name='pdbligand=MES:2-(N-MORPHOLINO)-ETHANESULFONIC+ACID'>MES</scene>, <scene name='pdbligand=SF4:IRON/SULFUR+CLUSTER'>SF4</scene></td></tr> | ||
<tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=4g9p FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=4g9p OCA], [https://pdbe.org/4g9p PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=4g9p RCSB], [https://www.ebi.ac.uk/pdbsum/4g9p PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=4g9p ProSAT]</span></td></tr> | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=4g9p FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=4g9p OCA], [https://pdbe.org/4g9p PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=4g9p RCSB], [https://www.ebi.ac.uk/pdbsum/4g9p PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=4g9p ProSAT]</span></td></tr> | ||
</table> | </table> | ||
== Function == | == Function == | ||
[https://www.uniprot.org/uniprot/ISPG_THET2 ISPG_THET2] | [https://www.uniprot.org/uniprot/ISPG_THET2 ISPG_THET2] | ||
| - | <div style="background-color:#fffaf0;"> | ||
| - | == Publication Abstract from PubMed == | ||
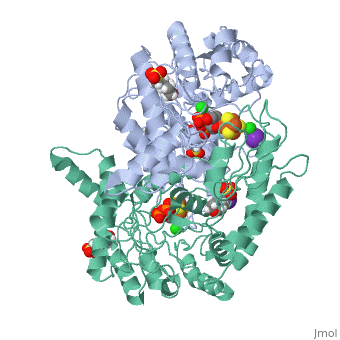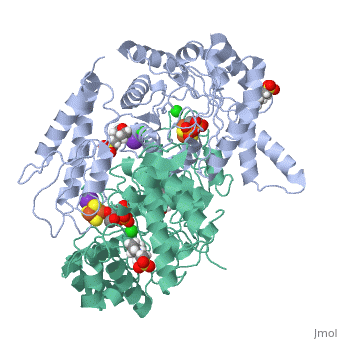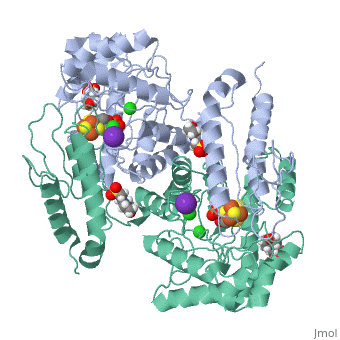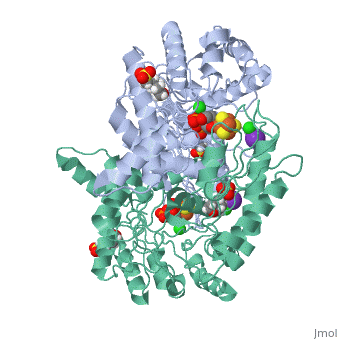
| - | Isoprenoid precursor biosynthesis occurs through the mevalonate or the methylerythritol phosphate (MEP) pathway, used i.e., by humans and by many human pathogens, respectively. In the MEP pathway, 2-C-methyl-d-erythritol-2,4-cyclo-diphosphate (MEcPP) is converted to (E)-1-hydroxy-2-methyl-but-2-enyl-4-diphosphate (HMBPP) by the iron-sulfur cluster enzyme HMBPP synthase (GcpE). The presented X-ray structure of the GcpE-MEcPP complex from Thermus thermophilus at 1.55A resolution provides valuable information about the catalytic mechanism and for rational inhibitor design. MEcPP binding inside the TIM-barrel funnel induces a 60 degrees rotation of the [4Fe-4S] cluster containing domain onto the TIM-barrel entrance. The apical iron of the [4Fe-4S] cluster ligates with the C3 oxygen atom of MEcPP. | ||
| - | |||
| - | Structure of the GcpE (IspG)-MEcPP complex from Thermus thermophilus.,Rekittke I, Jomaa H, Ermler U FEBS Lett. 2012 Sep 21;586(19):3452-7. doi: 10.1016/j.febslet.2012.07.070. Epub, 2012 Aug 9. PMID:22967895<ref>PMID:22967895</ref> | ||
| - | |||
| - | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| - | </div> | ||
| - | <div class="pdbe-citations 4g9p" style="background-color:#fffaf0;"></div> | ||
==See Also== | ==See Also== | ||
*[[IspG|IspG]] | *[[IspG|IspG]] | ||
| - | == References == | ||
| - | <references/> | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
Current revision
Structure of the GcpE-MEcPP (IspG) complex from Thermus thermophilus
| |||||||||||