We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Sandbox Reserved 1783
From Proteopedia
(Difference between revisions)
| Line 14: | Line 14: | ||
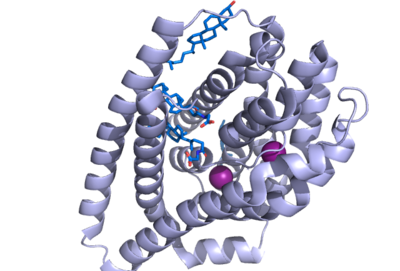
There are two significant areas in the NTCP structure that facilitate ligand binding, which are referred to as "patches." Residues 84-87 of NTCP are patch 1, which are located on the TM2-TM3 loop in the core domain. This is also considered the extracellular region of NTCP “tunnel." Residues 157-165 NTCP are associated with patch 2. They are located on N-terminal half of the TM5 in the panel domain (residue sequence: KGIVISLVL). Patch 2 is also located in th extracellular region. These residues' importance was determined through mutations of these residues and examined through pull-down assays (Asami, et. al, 2022). | There are two significant areas in the NTCP structure that facilitate ligand binding, which are referred to as "patches." Residues 84-87 of NTCP are patch 1, which are located on the TM2-TM3 loop in the core domain. This is also considered the extracellular region of NTCP “tunnel." Residues 157-165 NTCP are associated with patch 2. They are located on N-terminal half of the TM5 in the panel domain (residue sequence: KGIVISLVL). Patch 2 is also located in th extracellular region. These residues' importance was determined through mutations of these residues and examined through pull-down assays (Asami, et. al, 2022). | ||
| - | |||
| - | [[Image:Ntcp baby.png|400 px|right|thumb|Figure 1: NTCP separated by domains, a panel domain and a core domain involved in a conformational change]] | ||
| - | <scene name='95/952711/Ntcp_regions_color_coded/1'>NTCP Separated by Domains</scene> | ||
| - | |||
=== Binding Pocket === | === Binding Pocket === | ||
Revision as of 20:27, 20 March 2023
| This Sandbox is Reserved from February 27 through August 31, 2023 for use in the course CH462 Biochemistry II taught by R. Jeremy Johnson at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1765 through Sandbox Reserved 1795. |
To get started:
More help: Help:Editing |
Sodium Taurocholate Co-Transporting Peptide
| |||||||||||
References
- ↑ Goutam, K., Ielasi, F.S., Pardon, E. et al. Structural basis of sodium-dependent bile salt uptake into the liver. Nature 606, 1015–1020 (2022). DOI: 10.1038/s41586-022-04723-z.
- ↑ Maldonado-Valderrama, J., Wilde, P., Macierzanka, A., & Mackie, A. (2011). The role of bile salts in digestion. Advances in colloid and interface science, 165(1), 36–46. DOI: 10.1016/j.cis.2010.12.002.
- ↑ Asami J, Kimura KT, Fujita-Fujiharu Y, Ishida H, Zhang Z, Nomura Y, Liu K, Uemura T, Sato Y, Ono M, Yamamoto M, Noda T, Shigematsu H, Drew D, Iwata S, Shimizu T, Nomura N, Ohto U. Structure of the bile acid transporter and HBV receptor NTCP. Nature. 2022 Jun; 606 (7916):1021-1026. DOI: 10.1038/s41586-022-04845-4.