This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1786
From Proteopedia
(Difference between revisions)
| Line 12: | Line 12: | ||
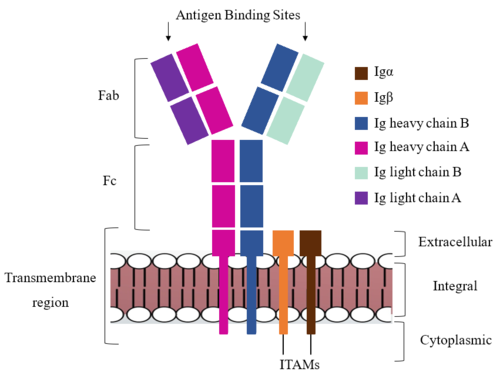
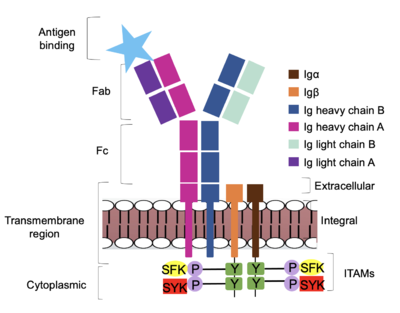
[[Image:IgM_structure_overview_diagram.png|500 px|right|thumb|'''Figure 1. IgM BCR Structure Overview.''' Depiction of the IgM BCR expressed on the membrane of a B cell. Includes all major components including the α/β heterodimer, heavy and light chains, antigen binding sites, and the ITAM region for signal transduction.]] | [[Image:IgM_structure_overview_diagram.png|500 px|right|thumb|'''Figure 1. IgM BCR Structure Overview.''' Depiction of the IgM BCR expressed on the membrane of a B cell. Includes all major components including the α/β heterodimer, heavy and light chains, antigen binding sites, and the ITAM region for signal transduction.]] | ||
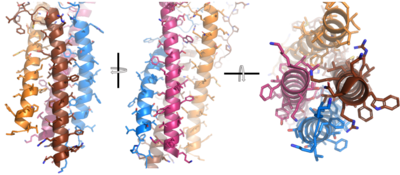
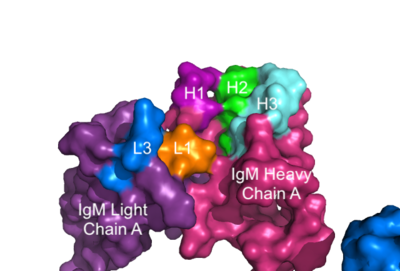
| - | + | The IgM BCR consists of six separate chains (Figure 1) that make up the three main [https://en.wikipedia.org/wiki/Protein_domain domains] typically found in a [https://bioxcell.com/educational-articles/antibody-structure/ general antibody structure]. A depiction of the IgM <scene name='95/952714/Colored_by_domain/6'>colored by domain</scene> shows two heavy and two light chains that together form the <b><span class="text-cyan">Fab region</span></b>, or variable fragment at the top of the molecule where the antigen binding sites are located. The two heavy chains extend below the <b><span class="text-cyan">Fab region</span></b> through the <b><span class="text-purple">Fc region</span></b> and eventually connect to the Igα/β heterodimer to form the <b><span class="text-orange">transmembrane region</span></b> which anchors the overall complex to the B cell. These regions are also labeled in Figure 1. The overall structure, expression, and function of the IgM BCR is strongly influenced by the <b><span class="text-orange">transmembrane region</span></b> in which Ig α/β interactions as a heterodimer influence cell surface expression, receptor assembly, and effective signal transduction. <ref name="Tolar">Tolar P, Pierce SK. Unveiling the B cell receptor structure. Science. 2022 Aug 19;377(6608):819-820. doi: 10.1126/science.add8065. Epub 2022 Aug 18.[http://dx.doi.org/10.1126/science.add8065 DOI:10.1126/science.add8065</ref>, <ref name="Dylke">Dylke J, Lopes J, Dang-Lawson M, Machtaler S, Matsuuchi L. Role of the extracellular and transmembrane domain of Ig-alpha/beta in assembly of the B cell antigen receptor (BCR). Immunol Lett. 2007 Sep 15;112(1):47-57. doi: 10.1016/j.imlet.2007.06.005. Epub 2007 Jul 23. [http://dx.doi.org/10.1016/j.imlet.2007.06.005 DOI:10.1016/j.imlet.2007.06.005</ref>. In each domain, interactions between individual chains are important to understand the complex as a whole. All future 3D depictions showing these interactions ues the same PyMOL file [https://www.rcsb.org/structure/7XQ8 (7xq8)] and will be <scene name='95/952714/Colored_by_chain/12'>colored by chain</scene> as in Figure 1. | |
{{Clear}} | {{Clear}} | ||
Revision as of 00:05, 21 April 2023
Human B-cell Antigen Receptor: IgM BCR
| |||||||||||
References
- ↑ Sathe A, Cusick JK. Biochemistry, Immunoglobulin M. 2022 Dec 19. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2023 Jan–. PMID: 32310455. https://pubmed.ncbi.nlm.nih.gov/32310455/
- ↑ 2.0 2.1 Su Q, Chen M, Shi Y, Zhang X, Huang G, Huang B, Liu D, Liu Z, Shi Y. Cryo-EM structure of the human IgM B cell receptor. Science. 2022 Aug 19;377(6608):875-880. doi: 10.1126/science.abo3923. Epub 2022, Aug 18. PMID:35981043 doi:http://dx.doi.org/10.1126/science.abo3923
- ↑ 3.0 3.1 3.2 3.3 Ma X, Zhu Y, Dong, Chen Y, Wang S, Yang D, Ma Z, Zhang A, Zhang F, Guo C, Huang Z. Cryo-EM structures of two human B cell receptor isotypes. Science. 2022 Aug 19;377(6608):880-885. doi: 10.1126/science.abo3828. Epub 2022, Aug 18. PMID:35981028 doi:http://dx.doi.org/10.1126/science.abo3828
- ↑ 4.0 4.1 4.2 Tolar P, Pierce SK. Unveiling the B cell receptor structure. Science. 2022 Aug 19;377(6608):819-820. doi: 10.1126/science.add8065. Epub 2022 Aug 18.[http://dx.doi.org/10.1126/science.add8065 DOI:10.1126/science.add8065
- ↑ 5.0 5.1 5.2 Dylke J, Lopes J, Dang-Lawson M, Machtaler S, Matsuuchi L. Role of the extracellular and transmembrane domain of Ig-alpha/beta in assembly of the B cell antigen receptor (BCR). Immunol Lett. 2007 Sep 15;112(1):47-57. doi: 10.1016/j.imlet.2007.06.005. Epub 2007 Jul 23. [http://dx.doi.org/10.1016/j.imlet.2007.06.005 DOI:10.1016/j.imlet.2007.06.005
- ↑ Daniels R, Kurowski B, Johnson AE, Hebert DN. N-linked glycans direct the cotranslational folding pathway of influenza hemagglutinin. Mol Cell. 2003 Jan;11(1):79-90. PMID:12535523 doi:10.1016/s1097-2765(02)00821-3
- ↑ Bakshi T, Pham D, Kaur R, Sun B. Hidden Relationships between N-Glycosylation and Disulfide Bonds in Individual Proteins. Int J Mol Sci. 2022 Mar 29;23(7):3742. PMID:35409101 doi:10.3390/ijms23073742
- ↑ Mirazimi A, Svensson L. Carbohydrates facilitate correct disulfide bond formation and folding of rotavirus VP7. J Virol. 1998 May;72(5):3887-92. PMID:9557673 doi:10.1128/JVI.72.5.3887-3892.1998
- ↑ Zhou T, Georgiev I, Wu X, Yang ZY, Dai K, Finzi A, Do Kwon Y, Scheid JF, Shi W, Xu L, Yang Y, Zhu J, Nussenzweig MC, Sodroski J, Shapiro L, Nabel GJ, Mascola JR, Kwong PD. Structural basis for broad and potent neutralization of HIV-1 by antibody VRC01. Science. 2010 Aug 13;329(5993):811-7. Epub 2010 Jul 8. PMID:20616231 doi:10.1126/science.1192819
Student Contributors
DeTonyeá Dickson, Allison Goss, Jackson Payton