This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Ben Whiteside
From Proteopedia
(Difference between revisions)
| Line 20: | Line 20: | ||
====Amidated C-Terminus==== | ====Amidated C-Terminus==== | ||
The <scene name='10/1038819/Amidated_c_term/9'>C-Terminus</scene> of amylin contains an amide group, rather than a carboxylic acid group. This chemical modification allows for more extensive hydrogen bonding to nearby residues, due to the added hydrogen bond donor on the NH2 group. In turn, this allows for favorable hydrogen bonds between S129 of the transmembrane domain and the main chain of Y37 on amylin. This interaction causes a "kink" in the random coil of amylin, displacing Y37 into a hydrophobic pocket, allowing for favorable hydrophobic interactions with W79 of the transmembrane domain. This amidation is thought to be a post-translational modification. | The <scene name='10/1038819/Amidated_c_term/9'>C-Terminus</scene> of amylin contains an amide group, rather than a carboxylic acid group. This chemical modification allows for more extensive hydrogen bonding to nearby residues, due to the added hydrogen bond donor on the NH2 group. In turn, this allows for favorable hydrogen bonds between S129 of the transmembrane domain and the main chain of Y37 on amylin. This interaction causes a "kink" in the random coil of amylin, displacing Y37 into a hydrophobic pocket, allowing for favorable hydrophobic interactions with W79 of the transmembrane domain. This amidation is thought to be a post-translational modification. | ||
| - | === | + | === Two-Domain Model of Amylin Binding === |
| + | It is hypothesized that amylin binds to the receptor via a two-domain model. The model suggests a series of steps for how amylin binds. First, the c-terminus of amylin binds to the n terminus of the extracellular domain of the receptor. This binding factors the alignment of amylin's n-terminus to the primary GPCR binding site. This activates the GPCR, leading to subsequent activation of adenylyl cyclase and cAMP release. | ||
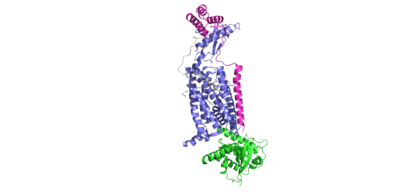
<scene name='10/1038828/Ramp_ctr_interface/9'>RAMP CTR Interface </scene> is a key interaction that stabilizes the protein complex and positions the receptor to favorably bind to amylin. The RAMP-CTR interface extends into the plasma membrane, providing additional non-covalent bonding between the protein complex and the cell membrane. | <scene name='10/1038828/Ramp_ctr_interface/9'>RAMP CTR Interface </scene> is a key interaction that stabilizes the protein complex and positions the receptor to favorably bind to amylin. The RAMP-CTR interface extends into the plasma membrane, providing additional non-covalent bonding between the protein complex and the cell membrane. | ||
==== Extracellular Domain - RAMP interactions ==== | ==== Extracellular Domain - RAMP interactions ==== | ||
Revision as of 17:54, 23 April 2024
AMYR
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Ransey E, Paredes E, Dey SK, Das SR, Heroux A, Macbeth MR. Crystal structure of the Entamoeba histolytica RNA lariat debranching enzyme EhDbr1 reveals a catalytic Zn(2+) /Mn(2+) heterobinucleation. FEBS Lett. 2017 Jul;591(13):2003-2010. doi: 10.1002/1873-3468.12677. Epub 2017, Jun 14. PMID:28504306 doi:http://dx.doi.org/10.1002/1873-3468.12677
- ↑ Cao J, Belousoff MJ, Liang YL, Johnson RM, Josephs TM, Fletcher MM, Christopoulos A, Hay DL, Danev R, Wootten D, Sexton PM. A structural basis for amylin receptor phenotype. Science. 2022 Mar 25;375(6587):eabm9609. PMID:35324283 doi:10.1126/science.abm9609
Student Contributors
Andrew Helmerich,Mathias Vander Eide, Ben Whiteside