This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox324
From Proteopedia
| Line 2: | Line 2: | ||
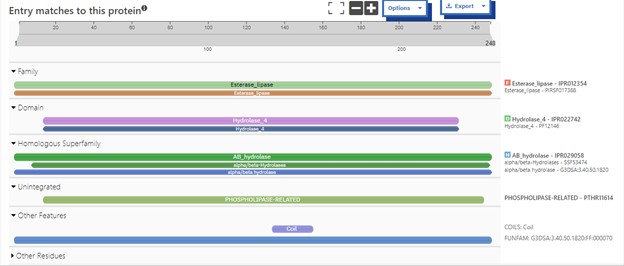
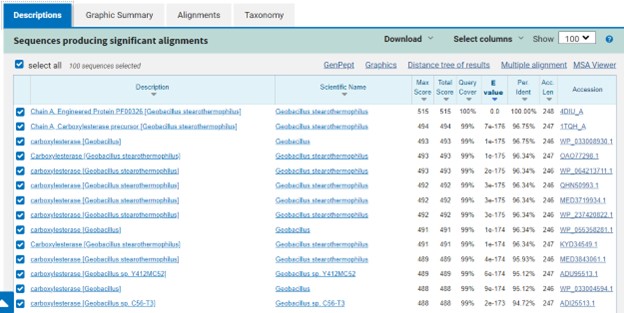
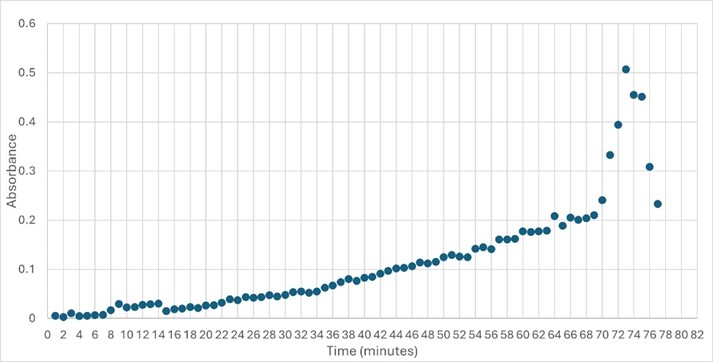
The protein 4DIU has been identified and characterized as a potential esterase enzyme based on in-depth analyses including bioinformatics, molecular docking studies, and laboratory experiments. Online tools such as BLAST, Dali, and InterPro often matched 4DIU with carboxylesterases and other esterase-like proteins. This suggests that its function involves hydrolysis of esters. In part to data from Swiss Dock, 4DIU is predicted to have binding sites with an affinity to substrates such as acetate, butyrate, phosphate, proline, decanoate, and dodecanoate. The protein 4DIU was grown, harvested, and isolated in a laboratory. SDS-PAGE analysis was used to confirm the identity of the protein via molecular weight. Once identified the enzyme's activity on the substrate p-nitrophenyl acetate was measured. This substrate was chosen because many hydrolases have an affinity for it and can hydrolyze the substrate. It was determined that the 4DIU protein functions best at a pH of 6. | The protein 4DIU has been identified and characterized as a potential esterase enzyme based on in-depth analyses including bioinformatics, molecular docking studies, and laboratory experiments. Online tools such as BLAST, Dali, and InterPro often matched 4DIU with carboxylesterases and other esterase-like proteins. This suggests that its function involves hydrolysis of esters. In part to data from Swiss Dock, 4DIU is predicted to have binding sites with an affinity to substrates such as acetate, butyrate, phosphate, proline, decanoate, and dodecanoate. The protein 4DIU was grown, harvested, and isolated in a laboratory. SDS-PAGE analysis was used to confirm the identity of the protein via molecular weight. Once identified the enzyme's activity on the substrate p-nitrophenyl acetate was measured. This substrate was chosen because many hydrolases have an affinity for it and can hydrolyze the substrate. It was determined that the 4DIU protein functions best at a pH of 6. | ||
| - | <StructureSection load=' | + | <StructureSection load='4DIU' size='340' side='right' caption='Structural Model of Protein 4DIU' scene=''> |
<ref>PMID:21638687</ref> | <ref>PMID:21638687</ref> | ||
Revision as of 23:04, 25 April 2024
Protein 4DIU: Structure, Function, and Significance in Biological Systems
The protein 4DIU has been identified and characterized as a potential esterase enzyme based on in-depth analyses including bioinformatics, molecular docking studies, and laboratory experiments. Online tools such as BLAST, Dali, and InterPro often matched 4DIU with carboxylesterases and other esterase-like proteins. This suggests that its function involves hydrolysis of esters. In part to data from Swiss Dock, 4DIU is predicted to have binding sites with an affinity to substrates such as acetate, butyrate, phosphate, proline, decanoate, and dodecanoate. The protein 4DIU was grown, harvested, and isolated in a laboratory. SDS-PAGE analysis was used to confirm the identity of the protein via molecular weight. Once identified the enzyme's activity on the substrate p-nitrophenyl acetate was measured. This substrate was chosen because many hydrolases have an affinity for it and can hydrolyze the substrate. It was determined that the 4DIU protein functions best at a pH of 6.
| |||||||||||
References
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ https://doi.org/10.1186%2F1471-2180-12-27