This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox324
From Proteopedia
(Difference between revisions)
| Line 46: | Line 46: | ||
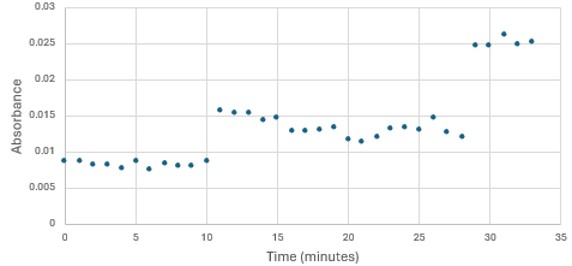
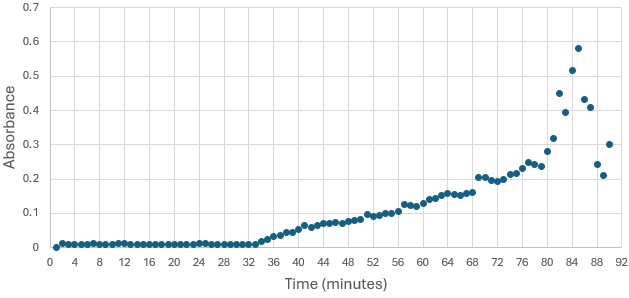
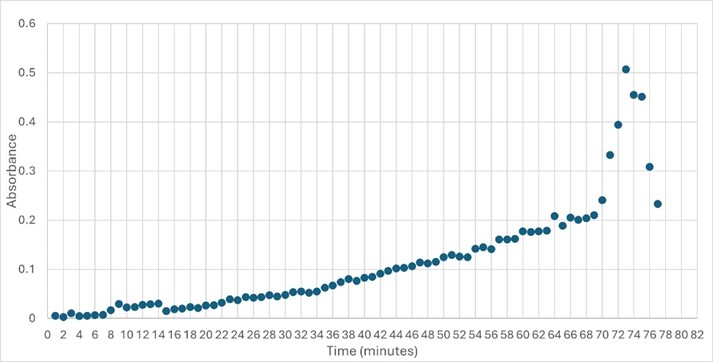
'''Figure 5: ''' Trial 2 of 4DIU enzymatic activity measured over time via change in absorbance at 405nm in pH 6 buffer | '''Figure 5: ''' Trial 2 of 4DIU enzymatic activity measured over time via change in absorbance at 405nm in pH 6 buffer | ||
| - | == Structural highlights of 4DIU == | + | == Structural highlights of 4DIU ==<ref>PMID:21638687</ref> |
| - | + | ||
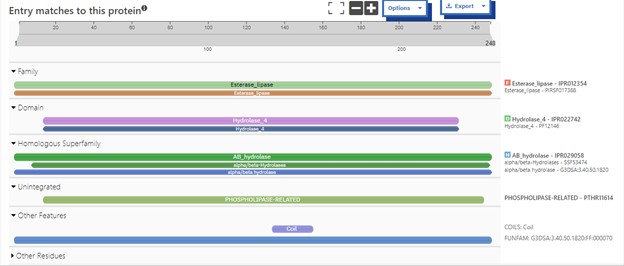
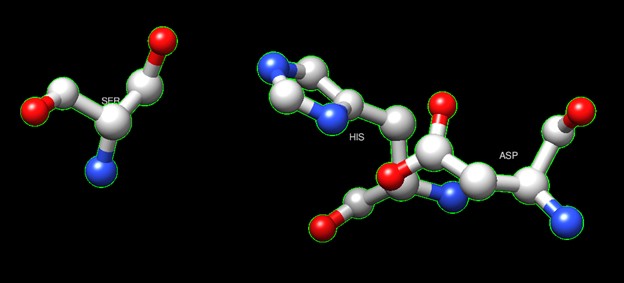
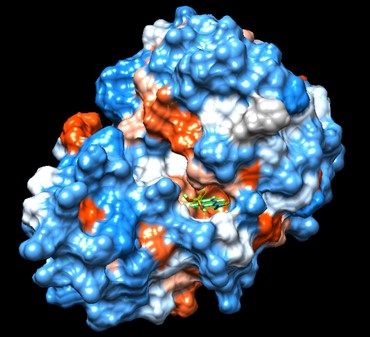
One of the distinctive structural characteristics of this protein is the alpha/beta-hydrolase (ABH) fold. Evidence for this fold consists of its structure (an open β-sheet surrounded by α-helices) and the presence of what is known as the "catalytic triad" (consisting of a nucleophile, an acid, and histidine)<ref>Holmquist, M. Alpha Beta-Hydrolase Fold Enzymes Structures, Functions and Mechanisms. Current Protein and Peptide Science 2000, 1 (2), 209–235. https://doi.org/10.2174/1389203003381405.</ref>. The presence of the active site residues His A 222, Asp A 192, and Ser A 93, as determined by SPRITE, Chimera, etc., confirms the presence of this catalytic triad. | One of the distinctive structural characteristics of this protein is the alpha/beta-hydrolase (ABH) fold. Evidence for this fold consists of its structure (an open β-sheet surrounded by α-helices) and the presence of what is known as the "catalytic triad" (consisting of a nucleophile, an acid, and histidine)<ref>Holmquist, M. Alpha Beta-Hydrolase Fold Enzymes Structures, Functions and Mechanisms. Current Protein and Peptide Science 2000, 1 (2), 209–235. https://doi.org/10.2174/1389203003381405.</ref>. The presence of the active site residues His A 222, Asp A 192, and Ser A 93, as determined by SPRITE, Chimera, etc., confirms the presence of this catalytic triad. | ||
Revision as of 16:32, 29 April 2024
| |||||||||||
References
- ↑ Fukami, T.; Yokoi, T. The Emerging Role of Human Esterases. Drug Metabolism and Pharmacokinetics 2012, 27 (5), 466–477. https://doi.org/10.2133/dmpk.dmpk-12-rv-042.
- ↑ Tokudome, Y.; Katayanagi, M.; Hashimoto, F. Esterase Activity and Intracellular Localization in Reconstructed Human Epidermal Cultured Skin Models. Annals of Dermatology 2015, 27 (3), 269. https://doi.org/10.5021/ad.2015.27.3.269.
- ↑ Williams, F. M. Clinical Significance of Esterases in Man. Clinical pharmacokinetics 1985, 10 (5), 392–403. https://doi.org/10.2165/00003088-198510050-00002.
- ↑ Zhang, S.; Sun, W.; Xu, L.; Zheng, X.; Chu, X.; Tian, J.; Wu, N.; Fan, Y. Identification of the Para-Nitrophenol Catabolic Pathway, and Characterization of Three Enzymes Involved in the Hydroquinone Pathway, in Pseudomonas Sp. 1-7. BMC Microbiology 2012, 12 (1). https://doi.org/10.1186/1471-2180-12-27.
- ↑ Vázquez-Mayorga, E.; Díaz-Sánchez, Á.; Dagda, R.; Domínguez-Solís, C.; Dagda, R.; Coronado-Ramírez, C.; Martínez-Martínez, A. Novel Redox-Dependent Esterase Activity (EC 3.1.1.2) for DJ-1: Implications for Parkinson’s Disease. International Journal of Molecular Sciences 2016, 17 (8), 1346. https://doi.org/10.3390/ijms17081346.
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Holmquist, M. Alpha Beta-Hydrolase Fold Enzymes Structures, Functions and Mechanisms. Current Protein and Peptide Science 2000, 1 (2), 209–235. https://doi.org/10.2174/1389203003381405.