We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Sandbox Reserved 1846
From Proteopedia
(Difference between revisions)
| Line 23: | Line 23: | ||
=== Catalytic Triad === | === Catalytic Triad === | ||
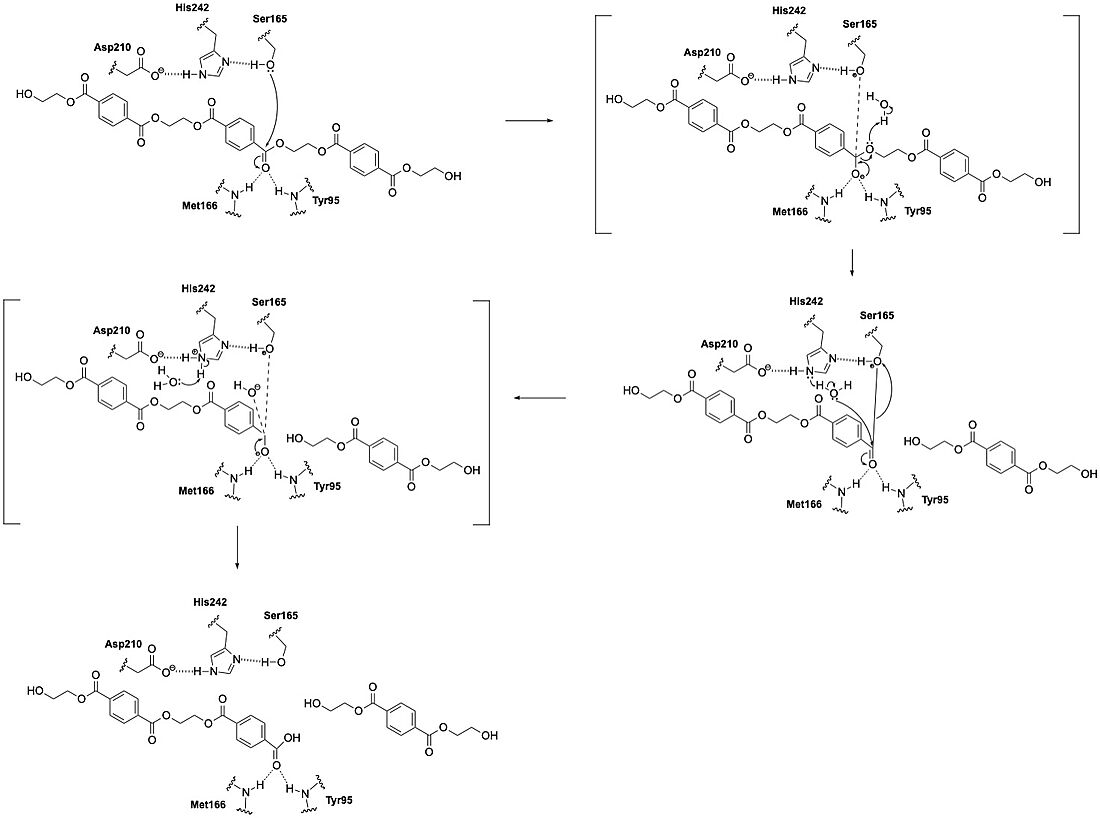
LCC catalyzes the breakdown of PET using a classic serine hydrolase mechanism involving a <scene name='10/1075246/Catalytic_triad3/1'>catalytic triad</scene> of Ser165, His242, and Asp210. The reaction begins when His242 deprotonates Ser165, which activates it as a nucleophile. Ser165 then attacks the carbonyl carbon of an ester bond in the PET polymer. This forms a tetrahedral intermediate. This intermediate is stabilized by an oxyanion hole. The oxyanion hole is formed by the backbone amides of Met166 and Tyr95. The intermediate collapses; one product is released and an acyl-enzyme intermediate is formed. A water molecule, activated by His242, then attacks the acyl-enzyme, releasing the second product and resetting the enzyme’s active site. | LCC catalyzes the breakdown of PET using a classic serine hydrolase mechanism involving a <scene name='10/1075246/Catalytic_triad3/1'>catalytic triad</scene> of Ser165, His242, and Asp210. The reaction begins when His242 deprotonates Ser165, which activates it as a nucleophile. Ser165 then attacks the carbonyl carbon of an ester bond in the PET polymer. This forms a tetrahedral intermediate. This intermediate is stabilized by an oxyanion hole. The oxyanion hole is formed by the backbone amides of Met166 and Tyr95. The intermediate collapses; one product is released and an acyl-enzyme intermediate is formed. A water molecule, activated by His242, then attacks the acyl-enzyme, releasing the second product and resetting the enzyme’s active site. | ||
| - | [[Image:Mechanism_(1).jpeg|1100 px]] | + | [[Image:Mechanism_(1).jpeg|1100 px|left|Figure 1: Mechanism]] |
=== Ligand Binding Pocket === | === Ligand Binding Pocket === | ||
The <scene name='10/1075246/4eb0_with_colored_ligand_stick/1'>substrate-binding site</scene> of LCC is a long, mainly hydrophobic groove that accommodates PET chains. This groove includes three subsites—designated −2, −1, and +1—that interact with specific PET units near the scissile ester bond. Hydrophobic residues such as F125, V212, M166, and F243 line the groove and facilitate binding by interacting with the aromatic rings of the PET molecule. These interactions help align the substrate in the correct position for catalysis. | The <scene name='10/1075246/4eb0_with_colored_ligand_stick/1'>substrate-binding site</scene> of LCC is a long, mainly hydrophobic groove that accommodates PET chains. This groove includes three subsites—designated −2, −1, and +1—that interact with specific PET units near the scissile ester bond. Hydrophobic residues such as F125, V212, M166, and F243 line the groove and facilitate binding by interacting with the aromatic rings of the PET molecule. These interactions help align the substrate in the correct position for catalysis. | ||
| - | [[Image:Final rayed image of binding pocket.png|400 px|right|thumb|Figure | + | [[Image:Final rayed image of binding pocket.png|400 px|right|thumb|Figure 2:Ser,His,Asp Catalytic Triad]] |
<scene name='10/1075246/4eb0_surface_w_stick_ligand/1'>Molecular surface view of enzyme-ligand interaction</scene> | <scene name='10/1075246/4eb0_surface_w_stick_ligand/1'>Molecular surface view of enzyme-ligand interaction</scene> | ||
| Line 64: | Line 64: | ||
N246M | N246M | ||
| - | The asparagine side chain is mutated to aspartic acid and methionine to increase thermostability. The wild-type protein has a melting point of 84.7°C. The N246D mutation (asparagine to aspartic acid) replaces a polar neutral side chain with a negatively charged one, which potentially increases electrostatic interactions or salt bridges that stabilize the protein. This results in a melting point of 87.9°C. The N246M mutation (asparagine to methionine) introduces a bulkier, hydrophobic side chain. This increases the internal packing of the protein core. This mutant has a melting point of 88.0°C | + | The asparagine side chain is mutated to aspartic acid and methionine to increase thermostability. The wild-type protein has a melting point of 84.7°C. The N246D mutation (asparagine to aspartic acid) replaces a polar neutral side chain with a negatively charged one, which potentially increases electrostatic interactions or salt bridges that stabilize the protein. This results in a melting point of 87.9°C. The N246M mutation (asparagine to methionine) introduces a bulkier, hydrophobic side chain. This increases the internal packing of the protein core. This mutant has a melting point of 88.0°C. |
=== S283 & D238 === | === S283 & D238 === | ||
Revision as of 18:42, 10 April 2025
| This Sandbox is Reserved from March 18 through September 1, 2025 for use in the course CH462 Biochemistry II taught by R. Jeremy Johnson and Mark Macbeth at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1828 through Sandbox Reserved 1846. |
To get started:
More help: Help:Editing |
Leaf Branch Compost Cutinase
| |||||||||||
References
- ↑ 1.0 1.1 Tournier V, Topham CM, Gilles A, David B, Folgoas C, Moya-Leclair E, Kamionka E, Desrousseaux ML, Texier H, Gavalda S, Cot M, Guemard E, Dalibey M, Nomme J, Cioci G, Barbe S, Chateau M, Andre I, Duquesne S, Marty A. An engineered PET depolymerase to break down and recycle plastic bottles. Nature. 2020 Apr;580(7802):216-219. doi: 10.1038/s41586-020-2149-4. Epub 2020 Apr, 8. PMID:32269349 doi:http://dx.doi.org/10.1038/s41586-020-2149-4
- ↑ Kolattukudy PE. Biopolyester membranes of plants: cutin and suberin. Science. 1980 May 30;208(4447):990-1000. PMID:17779010 doi:10.1126/science.208.4447.990
- ↑ Burgin T, Pollard BC, Knott BC, Mayes HB, Crowley MF, McGeehan JE, Beckham GT, Woodcock HL. The reaction mechanism of the Ideonella sakaiensis PETase enzyme. Commun Chem. 2024 Mar 27;7(1):65. PMID:38538850 doi:10.1038/s42004-024-01154-x
- ↑ Landrigan PJ, Stegeman JJ, Fleming LE, Allemand D, Anderson DM, Backer LC, Brucker-Davis F, Chevalier N, Corra L, Czerucka D, Bottein MD, Demeneix B, Depledge M, Deheyn DD, Dorman CJ, Fénichel P, Fisher S, Gaill F, Galgani F, Gaze WH, Giuliano L, Grandjean P, Hahn ME, Hamdoun A, Hess P, Judson B, Laborde A, McGlade J, Mu J, Mustapha A, Neira M, Noble RT, Pedrotti ML, Reddy C, Rocklöv J, Scharler UM, Shanmugam H, Taghian G, van de Water JAJM, Vezzulli L, Weihe P, Zeka A, Raps H, Rampal P. Human Health and Ocean Pollution. Ann Glob Health. 2020 Dec 3;86(1):151. PMID:33354517 doi:10.5334/aogh.2831
- ↑ Jambeck JR, Geyer R, Wilcox C, Siegler TR, Perryman M, Andrady A, Narayan R, Law KL. Marine pollution. Plastic waste inputs from land into the ocean. Science. 2015 Feb 13;347(6223):768-71. PMID:25678662 doi:10.1126/science.1260352
Student Contributors
Ashley Callaghan Rebecca Hoff Simone McCowan