We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Sandbox Reserved 1849
From Proteopedia
(Difference between revisions)
| Line 14: | Line 14: | ||
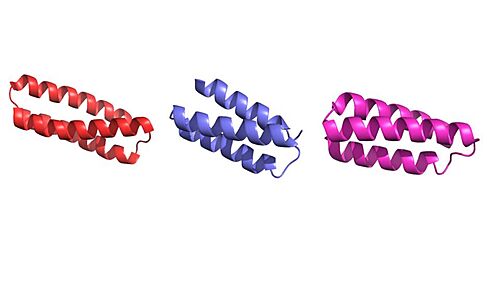
The primary goal of the mini binders is to prevent the spike proteins from binding to ACE2, and when the mini binders are bound to the spike protein, the virus is unable to anchor itself to the host protein <ref name="Longxing">PMID:32907861</ref>. Because the mini binders have a greater binding affinity than ACE2 for the spike protein, they are able to effectively prevent the entry of the virus and ultimately prevent an immune response <ref name="Longxing">PMID:32907861</ref>. Targeting this specific interaction between the COVID-19 spike protein has proven effective and is hopeful target for future therapeutics to treat the virus <ref name="Huang">PMID:32747721</ref>. LCB1 proved to be quite effective at weakening the immune response, compared to the other mini binders, which can be explained by the binding interface between the spike protein and LCB1 <ref name="Longxing">PMID:32907861</ref>. | The primary goal of the mini binders is to prevent the spike proteins from binding to ACE2, and when the mini binders are bound to the spike protein, the virus is unable to anchor itself to the host protein <ref name="Longxing">PMID:32907861</ref>. Because the mini binders have a greater binding affinity than ACE2 for the spike protein, they are able to effectively prevent the entry of the virus and ultimately prevent an immune response <ref name="Longxing">PMID:32907861</ref>. Targeting this specific interaction between the COVID-19 spike protein has proven effective and is hopeful target for future therapeutics to treat the virus <ref name="Huang">PMID:32747721</ref>. LCB1 proved to be quite effective at weakening the immune response, compared to the other mini binders, which can be explained by the binding interface between the spike protein and LCB1 <ref name="Longxing">PMID:32907861</ref>. | ||
| - | + | ==Expectations of this page== | |
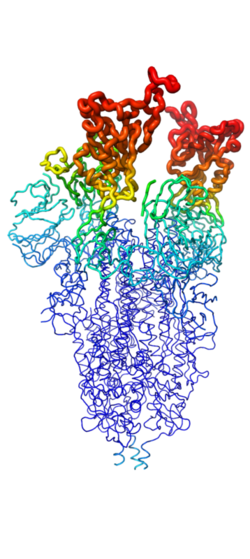
This page will dive into the method of mini binders effectively preventing the entry of the viral SARS-CoV-2 spike protein to enter a host cell. Taking a closer look at the structure of the SARS-CoV-2 spike protein, this will provide a deeper understanding of how it binds to host a cell membrane (ACE2) and give context to how the mini binders are capable of binding to it. Understanding the ACE2 receptor on host cells will give an expectation for how mini binders will bind to the spike protein, stealing the parts of the structure of ACE2 to create a protein with a greater binding affinity to outcompete ACE2. | This page will dive into the method of mini binders effectively preventing the entry of the viral SARS-CoV-2 spike protein to enter a host cell. Taking a closer look at the structure of the SARS-CoV-2 spike protein, this will provide a deeper understanding of how it binds to host a cell membrane (ACE2) and give context to how the mini binders are capable of binding to it. Understanding the ACE2 receptor on host cells will give an expectation for how mini binders will bind to the spike protein, stealing the parts of the structure of ACE2 to create a protein with a greater binding affinity to outcompete ACE2. | ||
Revision as of 13:50, 15 April 2025
| This Sandbox is Reserved from March 18 through September 1, 2025 for use in the course CH462 Biochemistry II taught by R. Jeremy Johnson and Mark Macbeth at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1828 through Sandbox Reserved 1846. |
To get started:
More help: Help:Editing |
SARS-COV2 Minibinders
| |||||||||||
Color Key
■ -> ACE2
■ -> Spike RBD
■ -> AHB2
■ -> LCB1
■ -> LCB3
References
[1] [2] [3] [4] [6] [5] [7] [8] [9]
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 1.17 1.18 1.19 1.20 1.21 Cao L, Goreshnik I, Coventry B, Case JB, Miller L, Kozodoy L, Chen RE, Carter L, Walls AC, Park YJ, Strauch EM, Stewart L, Diamond MS, Veesler D, Baker D. De novo design of picomolar SARS-CoV-2 miniprotein inhibitors. Science. 2020 Oct 23;370(6515):426-431. PMID:32907861 doi:10.1126/science.abd9909
- ↑ 2.0 2.1 2.2 2.3 2.4 Case JB, Chen RE, Cao L, Ying B, Winkler ES, Johnson M, Goreshnik I, Pham MN, Shrihari S, Kafai NM, Bailey AL, Xie X, Shi PY, Ravichandran R, Carter L, Stewart L, Baker D, Diamond MS. Ultrapotent miniproteins targeting the SARS-CoV-2 receptor-binding domain protect against infection and disease. Cell Host Microbe. 2021 Jul 14;29(7):1151-1161.e5. PMID:34192518 doi:10.1016/j.chom.2021.06.008
- ↑ 3.0 3.1 Sang P, Chen YQ, Liu MT, Wang YT, Yue T, Li Y, Yin YR, Yang LQ. Electrostatic Interactions Are the Primary Determinant of the Binding Affinity of SARS-CoV-2 Spike RBD to ACE2: A Computational Case Study of Omicron Variants. Int J Mol Sci. 2022 Nov 26;23(23):14796. PMID:36499120 doi:10.3390/ijms232314796
- ↑ 4.00 4.01 4.02 4.03 4.04 4.05 4.06 4.07 4.08 4.09 4.10 4.11 4.12 Huang Y, Yang C, Xu XF, Xu W, Liu SW. Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacol Sin. 2020 Sep;41(9):1141-1149. doi: 10.1038/s41401-020-0485-4., Epub 2020 Aug 3. PMID:32747721 doi:http://dx.doi.org/10.1038/s41401-020-0485-4
- ↑ 5.0 5.1 5.2 5.3 Zhang J, Xiao T, Cai Y, Chen B. Structure of SARS-CoV-2 spike protein. Curr Opin Virol. 2021 Oct;50:173-182. PMID:34534731 doi:10.1016/j.coviro.2021.08.010
- ↑ 6.0 6.1 6.2 Yuan Y, Cao D, Zhang Y, Ma J, Qi J, Wang Q, Lu G, Wu Y, Yan J, Shi Y, Zhang X, Gao GF. Cryo-EM structures of MERS-CoV and SARS-CoV spike glycoproteins reveal the dynamic receptor binding domains. Nat Commun. 2017 Apr 10;8:15092. doi: 10.1038/ncomms15092. PMID:28393837 doi:http://dx.doi.org/10.1038/ncomms15092
- ↑ 7.0 7.1 7.2 Kuba K, Yamaguchi T, Penninger JM. Angiotensin-Converting Enzyme 2 (ACE2) in the Pathogenesis of ARDS in COVID-19. Front Immunol. 2021 Dec 22;12:732690. PMID:35003058 doi:10.3389/fimmu.2021.732690
- ↑ 8.0 8.1 Kuba K, Imai Y, Rao S, Gao H, Guo F, Guan B, Huan Y, Yang P, Zhang Y, Deng W, Bao L, Zhang B, Liu G, Wang Z, Chappell M, Liu Y, Zheng D, Leibbrandt A, Wada T, Slutsky AS, Liu D, Qin C, Jiang C, Penninger JM. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus-induced lung injury. Nat Med. 2005 Aug;11(8):875-9. PMID:16007097 doi:10.1038/nm1267
- ↑ 9.0 9.1 Valetti F, Gilardi G. Improvement of biocatalysts for industrial and environmental purposes by saturation mutagenesis. Biomolecules. 2013 Oct 8;3(4):778-811. PMID:24970191 doi:10.3390/biom3040778