This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1v3v
From Proteopedia
(New page: 200px<br /><applet load="1v3v" size="450" color="white" frame="true" align="right" spinBox="true" caption="1v3v, resolution 2.0Å" /> '''Crystal structure of ...) |
|||
| Line 1: | Line 1: | ||
| - | [[Image:1v3v.gif|left|200px]]<br /><applet load="1v3v" size=" | + | [[Image:1v3v.gif|left|200px]]<br /><applet load="1v3v" size="350" color="white" frame="true" align="right" spinBox="true" |
caption="1v3v, resolution 2.0Å" /> | caption="1v3v, resolution 2.0Å" /> | ||
'''Crystal structure of leukotriene B4 12-hydroxydehydrogenase/15-oxo-prostaglandin 13-reductase complexed with NADP and 15-oxo-PGE2'''<br /> | '''Crystal structure of leukotriene B4 12-hydroxydehydrogenase/15-oxo-prostaglandin 13-reductase complexed with NADP and 15-oxo-PGE2'''<br /> | ||
==Overview== | ==Overview== | ||
| - | The bifunctional leukotriene B(4) | + | The bifunctional leukotriene B(4) 12-hydroxydehydrogenase/15-oxo-prostaglandin 13-reductase (LTB(4) 12-HD/PGR) is an essential enzyme for eicosanoid inactivation. It is involved in the metabolism of the E and F series of 15-oxo-prostaglandins (15-oxo-PGs), leukotriene B(4) (LTB(4)), and 15-oxo-lipoxin A(4) (15-oxo-LXA(4)). Some nonsteroidal anti-inflammatory drugs (NSAIDs), which primarily act as cyclooxygenase inhibitors also inhibit LTB(4) 12-HD/PGR activity. Here we report the crystal structure of the LTB(4) 12-HD/PGR, the binary complex structure with NADP(+), and the ternary complex structure with NADP(+) and 15-oxo-PGE(2). In the ternary complex, both in the crystalline form and in solution, the enolate anion intermediate accumulates as a brown chromophore. PGE(2) contains two chains, but only the omega-chain of 15-oxo-PGE(2) was defined in the electron density map in the ternary complex structure. The omega-chain was identified at the hydrophobic pore on the dimer interface. The structure showed that the 15-oxo group forms hydrogen bonds with the 2'-hydroxyl group of nicotine amide ribose of NADP(+) and a bound water molecule to stabilize the enolate intermediate during the reductase reaction. The electron-deficient C13 atom of the conjugated enolate may be directly attacked by a hydride from the NADPH nicotine amide in a stereospecific manner. The moderate recognition of 15-oxo-PGE(2) is consistent with a broad substrate specificity of LTB(4) 12-HD/PGR. The structure also implies that a Src homology domain 3 may interact with the left-handed proline-rich helix at the dimer interface and regulate LTB(4) 12-HD/PGR activity by disruption of the substrate binding pore to accommodate the omega-chain. |
==About this Structure== | ==About this Structure== | ||
| - | 1V3V is a [http://en.wikipedia.org/wiki/Single_protein Single protein] structure of sequence from [http://en.wikipedia.org/wiki/Cavia_porcellus Cavia porcellus] with CL, NAP and 5OP as [http://en.wikipedia.org/wiki/ligands ligands]. Active as [http://en.wikipedia.org/wiki/15-oxoprostaglandin_13-oxidase 15-oxoprostaglandin 13-oxidase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=1.3.1.48 1.3.1.48] Full crystallographic information is available from [http:// | + | 1V3V is a [http://en.wikipedia.org/wiki/Single_protein Single protein] structure of sequence from [http://en.wikipedia.org/wiki/Cavia_porcellus Cavia porcellus] with <scene name='pdbligand=CL:'>CL</scene>, <scene name='pdbligand=NAP:'>NAP</scene> and <scene name='pdbligand=5OP:'>5OP</scene> as [http://en.wikipedia.org/wiki/ligands ligands]. Active as [http://en.wikipedia.org/wiki/15-oxoprostaglandin_13-oxidase 15-oxoprostaglandin 13-oxidase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=1.3.1.48 1.3.1.48] Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1V3V OCA]. |
==Reference== | ==Reference== | ||
| Line 18: | Line 18: | ||
[[Category: Kumasaka, T.]] | [[Category: Kumasaka, T.]] | ||
[[Category: Miyano, M.]] | [[Category: Miyano, M.]] | ||
| - | [[Category: RSGI, RIKEN | + | [[Category: RSGI, RIKEN Structural Genomics/Proteomics Initiative.]] |
[[Category: Shimizu, T.]] | [[Category: Shimizu, T.]] | ||
[[Category: Sugahara, M.]] | [[Category: Sugahara, M.]] | ||
| Line 32: | Line 32: | ||
[[Category: structural genomics]] | [[Category: structural genomics]] | ||
| - | ''Page seeded by [http:// | + | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Thu Feb 21 15:31:13 2008'' |
Revision as of 13:31, 21 February 2008
|
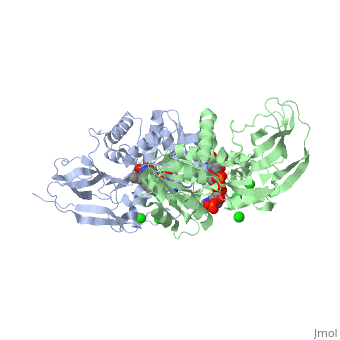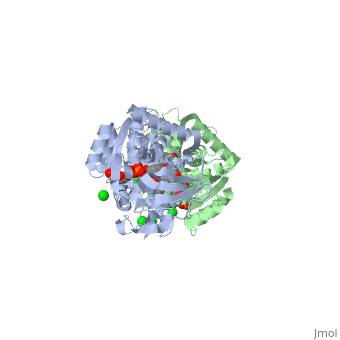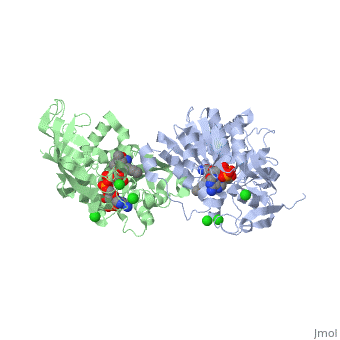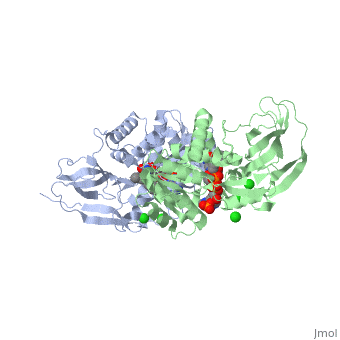
Crystal structure of leukotriene B4 12-hydroxydehydrogenase/15-oxo-prostaglandin 13-reductase complexed with NADP and 15-oxo-PGE2
Overview
The bifunctional leukotriene B(4) 12-hydroxydehydrogenase/15-oxo-prostaglandin 13-reductase (LTB(4) 12-HD/PGR) is an essential enzyme for eicosanoid inactivation. It is involved in the metabolism of the E and F series of 15-oxo-prostaglandins (15-oxo-PGs), leukotriene B(4) (LTB(4)), and 15-oxo-lipoxin A(4) (15-oxo-LXA(4)). Some nonsteroidal anti-inflammatory drugs (NSAIDs), which primarily act as cyclooxygenase inhibitors also inhibit LTB(4) 12-HD/PGR activity. Here we report the crystal structure of the LTB(4) 12-HD/PGR, the binary complex structure with NADP(+), and the ternary complex structure with NADP(+) and 15-oxo-PGE(2). In the ternary complex, both in the crystalline form and in solution, the enolate anion intermediate accumulates as a brown chromophore. PGE(2) contains two chains, but only the omega-chain of 15-oxo-PGE(2) was defined in the electron density map in the ternary complex structure. The omega-chain was identified at the hydrophobic pore on the dimer interface. The structure showed that the 15-oxo group forms hydrogen bonds with the 2'-hydroxyl group of nicotine amide ribose of NADP(+) and a bound water molecule to stabilize the enolate intermediate during the reductase reaction. The electron-deficient C13 atom of the conjugated enolate may be directly attacked by a hydride from the NADPH nicotine amide in a stereospecific manner. The moderate recognition of 15-oxo-PGE(2) is consistent with a broad substrate specificity of LTB(4) 12-HD/PGR. The structure also implies that a Src homology domain 3 may interact with the left-handed proline-rich helix at the dimer interface and regulate LTB(4) 12-HD/PGR activity by disruption of the substrate binding pore to accommodate the omega-chain.
About this Structure
1V3V is a Single protein structure of sequence from Cavia porcellus with , and as ligands. Active as 15-oxoprostaglandin 13-oxidase, with EC number 1.3.1.48 Full crystallographic information is available from OCA.
Reference
Structural basis of leukotriene B4 12-hydroxydehydrogenase/15-Oxo-prostaglandin 13-reductase catalytic mechanism and a possible Src homology 3 domain binding loop., Hori T, Yokomizo T, Ago H, Sugahara M, Ueno G, Yamamoto M, Kumasaka T, Shimizu T, Miyano M, J Biol Chem. 2004 May 21;279(21):22615-23. Epub 2004 Mar 8. PMID:15007077
Page seeded by OCA on Thu Feb 21 15:31:13 2008
Categories: 15-oxoprostaglandin 13-oxidase | Cavia porcellus | Single protein | Ago, H. | Hori, T. | Kumasaka, T. | Miyano, M. | RSGI, RIKEN Structural Genomics/Proteomics Initiative. | Shimizu, T. | Sugahara, M. | Ueno, G. | Yamamoto, M. | Yokomizo, T. | 5OP | CL | NAP | Riken structural genomics/proteomics initiative | Rossmann fold | Rsgi | Structural genomics