This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Tilman Schirmer/Sandbox 201
From Proteopedia
(→Intro) |
(→Substrate binding site) |
||
| Line 24: | Line 24: | ||
<br><br><br><br><br><br><br><br><br> | <br><br><br><br><br><br><br><br><br> | ||
| - | == | + | <applet load='2v0n' scene='User:Tilman_Schirmer/Sandbox_201/Substrate_binding_site/2' size='300' frame='true' align='right' caption='Diguanylate cyclase PleD (2v0n)' /> |
| - | <applet load='2v0n' scene='User:Tilman_Schirmer/Sandbox_201/Substrate_binding_site/2' size='300' frame='true' align='right' caption='2v0n' /> | ||
| - | <scene name='User:Tilman_Schirmer/Sandbox_201/Substrate_binding_site/3'> | + | The motif is part of the <scene name='User:Tilman_Schirmer/Sandbox_201/Substrate_binding_site/3'>substrate binding site</scene> as identified in the struture of the PleD - cene name='User:Tilman_Schirmer/Sandbox_201/Substrate_binding_ctp-a-s_mg/3'>GTP-α-S complex/Mg++</scene>. |
| - | <scene name='User:Tilman_Schirmer/Sandbox_201/Substrate_binding_ctp-a-s_mg/3'>Substrate binding site + ligand (CTP-alpha-S) + Mg++</scene> | ||
| - | + | ||
| + | |||
| + | <br><br><br><br><br><br><br><br><br><br><br><br><br> | ||
== Allosteric product binding site == | == Allosteric product binding site == | ||
Revision as of 20:20, 20 June 2009
PleD
Contents |
Intro
|
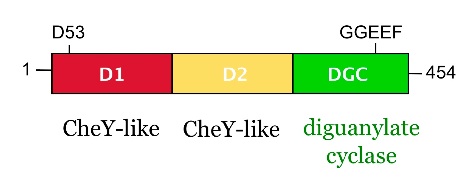
from Caulobacter crescentus is a response regulator with an unorthodox catalytic, diguanylate cyclase, output domain. It is composed of a canonical CheY-like response regulator receiver () domain,
a Rec-like () adaptor domain,
and a C-terminal domain that confers the catalytic acitvity.
The GGDEF domain is named after the highly conserved (in PleD it is GGEEF) that locates to a β-hairpin.
|
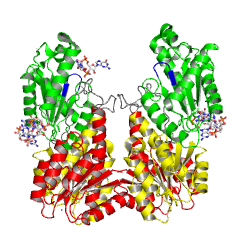
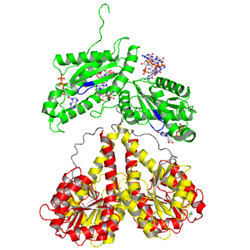
The motif is part of the as identified in the struture of the PleD - cene name='User:Tilman_Schirmer/Sandbox_201/Substrate_binding_ctp-a-s_mg/3'>GTP-α-S complex/Mg++</scene>.
Allosteric product binding site
|
C-di-GMP
Primary inhibition site (Ip)
Secondary inhibition site (Is)
Primary and secondary inhibition sites
Two conformations
|
|
Non-activated conformation
|
: , ,
Activated conformation
|