HMG-CoA Reductase
From Proteopedia
| Line 1: | Line 1: | ||
[[Image:1dqa opening.png|300px|left|thumb| Human HMG-CoA Reductase Catalytic Domain, [[1dqa]]]] | [[Image:1dqa opening.png|300px|left|thumb| Human HMG-CoA Reductase Catalytic Domain, [[1dqa]]]] | ||
{{STRUCTURE_1dq8| right| PDB=1dq8 | SCENE=HMG-CoA_Reductase/1dq8_starting_scene/1 |CAPTION= Crystal Structure of Human HMG-CoA Reductase Catalytic Domain, [[1dq8]] }} | {{STRUCTURE_1dq8| right| PDB=1dq8 | SCENE=HMG-CoA_Reductase/1dq8_starting_scene/1 |CAPTION= Crystal Structure of Human HMG-CoA Reductase Catalytic Domain, [[1dq8]] }} | ||
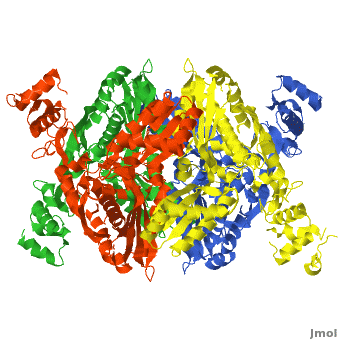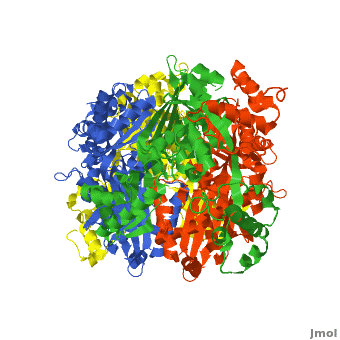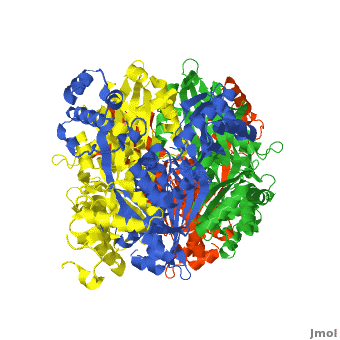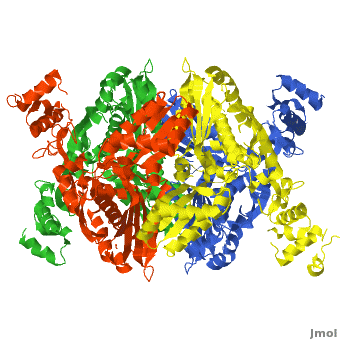
| - | [[HMG-CoA Reductase]] (or '''3-hydroxy-3-methyl-glutaryl-CoA reductasese''' or '''HMGR''') is the rate-controlling enzyme of the mevalonate pathway, responsible for cholesterol and other isoprenoid biosynthesis. HMGR is a | + | [[HMG-CoA Reductase]] (or '''3-hydroxy-3-methyl-glutaryl-CoA reductasese''' or '''HMGR''') is the rate-controlling enzyme of the mevalonate pathway, responsible for cholesterol and other isoprenoid biosynthesis. HMGR is a transmembrane protein, containing 8 domains, that is anchored in the membrane of the endoplasmic reticulum.<ref name="Roitelman">PMID:1374417</ref> It is the major target of the Statins, a cholesterol lowering drug class and the best selling pharmaceutical drugs in the world. |
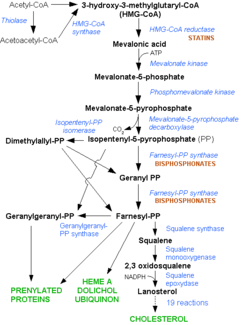
[[Image: HMG-CoA_reductase_pathway.png|250px|left|thumb| Mevalonate Pathway. Note the early stage at which the statins interfere in the pathway]] | [[Image: HMG-CoA_reductase_pathway.png|250px|left|thumb| Mevalonate Pathway. Note the early stage at which the statins interfere in the pathway]] | ||
| Line 14: | Line 14: | ||
<StructureSection load='1dq8' size='500' side='left' scene='HMG-CoA_Reductase/1dq8_starting_scene/1' caption='Crystal Structure of HMG-CoA'> | <StructureSection load='1dq8' size='500' side='left' scene='HMG-CoA_Reductase/1dq8_starting_scene/1' caption='Crystal Structure of HMG-CoA'> | ||
===General Structure=== | ===General Structure=== | ||
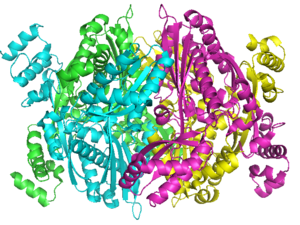
| - | There are two distinct classes of HMGRs, class I, which | + | There are two distinct classes of HMGRs, class I HMGRs, which are only found in eukaryotes and are membrane bound and class II HMGRs, which are found in prokaryotes and are soluble. <ref>PMID:11349148</ref> HMGR contains 8 transmembrane domains that have yet to be successfully crystallized, which anchor the protein to the membrane of the endoplasmic reticulum. <ref name="Roitelman"/> The catalytic portion of human HMGR forms a tetramer, with the individual monomers winding around each other. <ref name="Roitelman">PMID:1374417</ref> Within the tetramer, the monomers are arranged into <scene name='HMG-CoA_Reductase/1dq8_2_dimers/3'>two dimers</scene>, each of which contains <scene name='HMG-CoA_Reductase/1dq8_2_active_sites/2'>two active sites </scene>which are formed by residues form both monomers. Each monomer contains <scene name='HMG-CoA_Reductase/1dq8_star3_domains/2'>three domains </scene>, the <scene name='HMG-CoA_Reductase/1dq8_n_domain/2'>N-domain</scene>, the <scene name='HMG-CoA_Reductase/1dq8_l_domain/1'>L-Domain</scene>, and the <scene name='HMG-CoA_Reductase/1dq8_s_domain/1'>S-Domain</scene>. The L-domain is unique to HMGRs while the S-domain, which forms the binding site for NADP, resembles that of [[ferredoxin]]. The S and L domains are connected by a <scene name='HMG-CoA_Reductase/1dq8_cis_loop/6'>“cis-loop”</scene> which is essential for the HMG-binding site. <ref name="Roitelman"/> Salt bridges between residues R641 and E782 as well as <scene name='HMG-CoA_Reductase/1dq8_cis_loop/4'>hydrogen bonds</scene> between E700 and E700 on neighboring monomers compliment the largely hydrophobic dimer-dimer interface. <ref name="Roitelman"/> |
<br /> | <br /> | ||
Revision as of 07:46, 2 November 2010
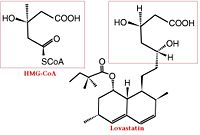
HMG-CoA Reductase (or 3-hydroxy-3-methyl-glutaryl-CoA reductasese or HMGR) is the rate-controlling enzyme of the mevalonate pathway, responsible for cholesterol and other isoprenoid biosynthesis. HMGR is a transmembrane protein, containing 8 domains, that is anchored in the membrane of the endoplasmic reticulum.[1] It is the major target of the Statins, a cholesterol lowering drug class and the best selling pharmaceutical drugs in the world.
Contents |
Biological Role
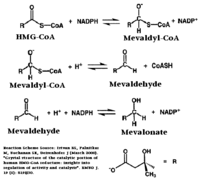
HMGR is among the most highly regulated enzymes in the human body. It catalyzes the formation of mevalonic acid, the committed step in the biosynthesis of sterols, most notably cholesterol. This reaction can be seen below where HMG-CoA is reduced by NADPH. Despite the poor reputation cholesterol has in the media, it is a critical component of cellular membranes as it is required to establish proper membrane permeability and fluidity. The mevalonate pathway is also responsible for synthesis of the oxygen transporting heme found in red blood cells. [2]
Structure
| |||||||||||
Medical Implications
| |||||||||||
Additional 3D Structures of HMG-CoA Reductase
HGMCR
1r7i - PmHMGCR catalytic domain - Pseudomonas mevalonii
HGMCR+statins
3cct, 3ccw, 3ccz, 3cd0, 3cd5, 3cd7, 3cda, 3cdb, 2r4f, 3bgl, 2q1l, 2q6b, 2q6c, 1hw8, 1hw9, 1hwi, 1hwj, 1hwk, 1hwl – HMG-CoA Reductase Catalytic Domain + Statins
1t02 - PmHMGCR catalytic domain + statin derivatives
HGMCR+cofactors
1r31 – PmHMGCR catalytic domain +CoA+mevalonate
1qax - PmHMGCR catalytic domain +HMG+CoA+NAD+
1qay - PmHMGCR catalytic domain +mevalonate+NAD+
1dq8 - hHMGCR catalytic domain (mutant) +CoA+HMG
1dq9 - hHMGCR catalytic domain (mutant)+HMG-CoA
1dqa - hHMGCR catalytic domain (mutant)+HMG+CoA+NADP+
Additional Resources
- See: Pharmaceutical Drug Targets For Additional Information about Drug Targets for Related Diseases
- See: Metabolic Disorders For Additional Information.
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 Roitelman J, Olender EH, Bar-Nun S, Dunn WA Jr, Simoni RD. Immunological evidence for eight spans in the membrane domain of 3-hydroxy-3-methylglutaryl coenzyme A reductase: implications for enzyme degradation in the endoplasmic reticulum. J Cell Biol. 1992 Jun;117(5):959-73. PMID:1374417
- ↑ 2.0 2.1 2.2 Meigs TE, Roseman DS, Simoni RD. Regulation of 3-hydroxy-3-methylglutaryl-coenzyme A reductase degradation by the nonsterol mevalonate metabolite farnesol in vivo. J Biol Chem. 1996 Apr 5;271(14):7916-22. PMID:8626470
- ↑ Istvan ES, Deisenhofer J. Structural mechanism for statin inhibition of HMG-CoA reductase. Science. 2001 May 11;292(5519):1160-4. PMID:11349148 doi:10.1126/science.1059344
- ↑ Song BL, Sever N, DeBose-Boyd RA. Gp78, a membrane-anchored ubiquitin ligase, associates with Insig-1 and couples sterol-regulated ubiquitination to degradation of HMG CoA reductase. Mol Cell. 2005 Sep 16;19(6):829-40. PMID:16168377 doi:10.1016/j.molcel.2005.08.009
- ↑ Goldstein JL, Brown MS. Regulation of the mevalonate pathway. Nature. 1990 Feb 1;343(6257):425-30. PMID:1967820 doi:http://dx.doi.org/10.1038/343425a0
- ↑ www.nhlbi.nih.gov/health/.../Diseases/.../CAD_WhatIs.html
- ↑ Endo A, Kuroda M, Tanzawa K. Competitive inhibition of 3-hydroxy-3-methylglutaryl coenzyme A reductase by ML-236A and ML-236B fungal metabolites, having hypocholesterolemic activity. FEBS Lett. 1976 Dec 31;72(2):323-6. PMID:16386050
- ↑ http://www.drugs.com/top200.html
- ↑ http://www.medicalnewstoday.com/articles/25046.php
- ↑ Istvan ES, Deisenhofer J. Structural mechanism for statin inhibition of HMG-CoA reductase. Science. 2001 May 11;292(5519):1160-4. PMID:11349148 doi:10.1126/science.1059344
- ↑ Corsini A, Maggi FM, Catapano AL. Pharmacology of competitive inhibitors of HMG-CoA reductase. Pharmacol Res. 1995 Jan;31(1):9-27. PMID:7784310
Proteopedia Page Contributors and Editors (what is this?)
David Canner, Michal Harel, Alexander Berchansky, Eran Hodis, Angel Herraez, Joel L. Sussman