Sandbox Reserved 321
From Proteopedia
| Line 31: | Line 31: | ||
==Substructure 2 of inhA== | ==Substructure 2 of inhA== | ||
| - | <scene name='Sandbox_Reserved_321/Substructure_2/1'>Substructure 2</scene> contains the c-terminal region of the molecule and consists of a small β strand (B-7), and two α helicies (A-6 and A-7) which are conected by a short five residue loop<ref name ="making drugs for inhA"/>. | + | <scene name='Sandbox_Reserved_321/Substructure_2/1'>Substructure 2</scene> contains the c-terminal region of the molecule and consists of a small β strand <scene name='Sandbox_Reserved_321/B-7/1'>(B-7)</scene>, and two α helicies <scene name='Sandbox_Reserved_321/A-6_and_a-7/1'>(A-6 and A-7)</scene> which are conected by a short five residue loop<ref name ="making drugs for inhA"/>. The C-terminal domain consits of two other α helicies <scene name='Sandbox_Reserved_321/A-8_and_a-9/1'>(A-8 and A-9)</scene><ref name ="making drugs for inhA"/>. |
=Physiological Function= | =Physiological Function= | ||
Revision as of 22:55, 31 March 2011
| This Sandbox is Reserved from January 10, 2010, through April 10, 2011 for use in BCMB 307-Proteins course taught by Andrea Gorrell at the University of Northern British Columbia, Prince George, BC, Canada. |
To get started:
More help: Help:Editing |
InhA
by Kelly Hrywkiw
| |||||||||
| 2h9i, resolution 2.20Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | |||||||||
| Gene: | inhA (Mycobacterium tuberculosis) | ||||||||
| Activity: | [acyl-carrier-protein_reductase_(NADH) Enoyl-[acyl-carrier-protein] reductase (NADH)], with EC number 1.3.1.9 | ||||||||
| Related: | 1zid | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
Introduction
The enzyme inhA is coded from the inhA gene that is simillar in sequence to the Salmonella typhimuriumgene which plays a role in fatty acid biosynthesis [1]. Inha is an NADH dependent trans enoyl-acyl ACP carrier protein that plays a role in the sysnthesis of Mycolic Acid, and is part of a short-chain dehydrogenase/reductase family [2][3]. Mycolic acids are long chain fatty acids that are essential in cell wall formation of the human pathogen Mycobacterium tuberculosisas well as other mycobateria such as Mycobacterium leprae[4]. Inha has been propsed as the target of the thionamide drugs, ethionamide (ETH) and isoniazid (INH), which have been used in treatment of mycobacterial infections [3].
Structure of inhA
|
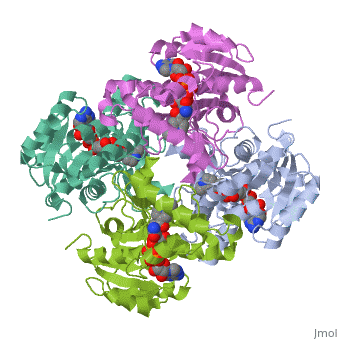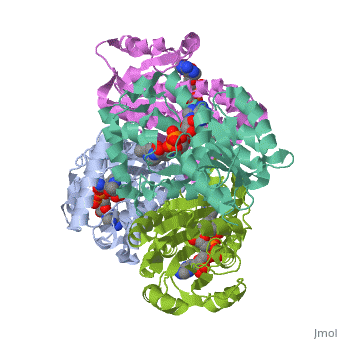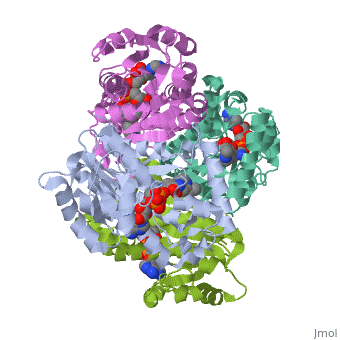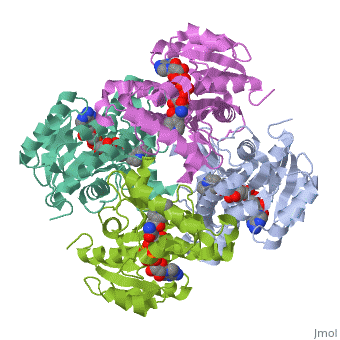
The overall strucuture of the inhA enzyme of Mycobacterium tuberculosisis a that of a homotetramer which is composed of a repeating subunit that consits of a single domain with two substructures that are connected by short loop[1][5].
Substructure 1 of inhA
consists of 6 parallel β strands and 4 α helices interwoven together to form a core α/β structure that contains the n-terminal domain[1]. The first substructure can be further broken down into two sections, the consisting of two β strands and two short α helicies [1]. The first section is connected to the by a β strand that crosses over the two domains, and leads into the second section initiating at the third α helix [1](A-3) is connected by a long loop to a 14 residue β strand that then leads into the fourth α helix [1]. A-4 then leads into a fifth strand β , followed by a 25 residue α helix , and into the final strand β [1].
Substructure 2 of inhA
contains the c-terminal region of the molecule and consists of a small β strand , and two α helicies which are conected by a short five residue loop[1]. The C-terminal domain consits of two other α helicies [1].
Physiological Function
Role in the Mycolic Acid Pathway
Protein Superfamilly
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 Sacchettini, James (New Rochelle, NY) 1999 INHA crystals and three dimensional structure United States Albert Einstein College of Medicine of Yeshiva University (Bronx, NY) 5882878 http://www.freepatentsonline.com/5882878.html
- ↑ Wang F, Langley R, Gulten G, Dover LG, Besra GS, Jacobs WR Jr, Sacchettini JC. Mechanism of thioamide drug action against tuberculosis and leprosy. J Exp Med. 2007 Jan 22;204(1):73-8. Epub 2007 Jan 16. PMID:17227913 doi:10.1084/jem.20062100
- ↑ 3.0 3.1 Molle V, Gulten G, Vilcheze C, Veyron-Churlet R, Zanella-Cleon I, Sacchettini JC, Jacobs WR Jr, Kremer L. Phosphorylation of InhA inhibits mycolic acid biosynthesis and growth of Mycobacterium tuberculosis. Mol Microbiol. 2010 Dec;78(6):1591-605. doi:, 10.1111/j.1365-2958.2010.07446.x. Epub 2010 Nov 9. PMID:21143326 doi:10.1111/j.1365-2958.2010.07446.x
- ↑ . PMID:216315890657
- ↑ Dias MV, Vasconcelos IB, Prado AM, Fadel V, Basso LA, de Azevedo WF Jr, Santos DS. Crystallographic studies on the binding of isonicotinyl-NAD adduct to wild-type and isoniazid resistant 2-trans-enoyl-ACP (CoA) reductase from Mycobacterium tuberculosis. J Struct Biol. 2007 Sep;159(3):369-80. Epub 2007 May 3. PMID:17588773 doi:http://dx.doi.org/10.1016/j.jsb.2007.04.009